Worm Breeder's Gazette 12(2): 75 (January 1, 1992)
These abstracts should not be cited in bibliographies. Material contained herein should be treated as personal communication and should be cited as such only with the consent of the author.
An efficient way to map the molecular location of a gene is to map the endpoints of duplications and/or deficiencies that terminate near that gene. These endpoints create restriction fragment length polymorphisms (RFLPs), which can be detected by comparing the genomes of non-duplication and duplication-bearing strains or non-deficiency and deficiency-bearing strains. Typically, this analysis is done using restriction enzymes that have 6bp recognition sequences (for example see Kramer et al. Cell 55, 555-565). Here, we describe a similar procedure that involves detecting duplication-specific polymorphisms using restriction enzymes with 8bp recognition sequences, which cut very infrequently in worm DNA. This method is less sensitive to small genomic polymorphisms, but enables large regions of the genome to be sampled. Thus, hundreds of kilobases can be rapidly scanned for the presence of endpoints. In addition, if there are rearrangements in the deficiency or duplication, this method easily detects them (see examples below). It has the disadvantage of requiring expensive enzymes and a pulsed field gel electrophoresis (PFGE) apparatus.
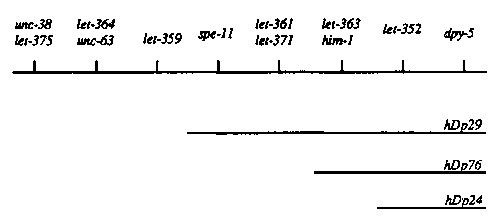
In our efforts to clone spe-11 ,which maps genetically between unc-38 and dpy-5 ,we have used this method to examine three duplications, hDp24 , hDp76 ,and hDp29 (provided by the Rose Lab), that terminate in this region. The genetic coverage of these duplications is as follows (reprinted from WBG I 1, #3, p.54 McKim, Rosenbluth, and Rose):
[See Figure 1]
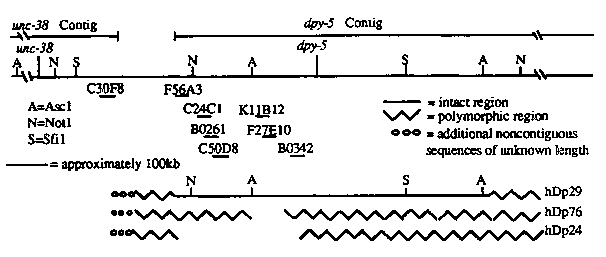
To prepare intact high molecular weight DNA for this analysis, we embedded L1 sin low melting temperature agarose and digested the agarose plugs with proteinase K. After extensive washes to remove the proteinase K, the plugs were digested with AscI, NotI, SfiI, or combinations of these enzymes. The digested DNA was run on pulsed field gels. Separation was achieved between approximately 10kb and 1Mb. The gels were blotted and probed with cosmids (provided by Alan Coulson and John Sulston) between unc-38 and dpy-5 .The polymorphism analysis was done by comparing the same strain with and without the duplication. Our map of this region, along with the position of mapped cosmids, is shown below:
[See Figure 2]
RFLP mapping of hDp29 suggests that this duplication ends between C3 OF8and F56A3 ,and that it is intact between the middle NotI and rightward AscI sites on the diagram above, but polymorphic outside of this region. Using F56A3 we detect a NotI polymorphism. However, we see no polymorphism with C3 OF8,suggesting that the duplication does not extend that far. C24C1 detects a different NotI polymorphism from F56A3 .However, a double digest with SfiI and NotI probed with C24C1 shows no polymorphism. In addition, C50D8 and K11B12 do not detect polymorphisms after double digesting the duplication strain DNA with NotI and SfiI or NotI and AscI. Thus, the duplication is intact between the middle NotI and rightward AscI sites but polymorphic outside of that region.
RFLP mapping of hDp24 reveals a large deletion and possibly an insertion. F56A3 detects a NotI polymorphism but C3 OF8does not, suggesting the duplication ends between these cosmids. The same size NotI polymorphism is detected with B0342 as with F56A3 .However, no polymorphisms are detected with C24C1 , B0261 , C5 OD8, K11B12 or F27E10 .Thus, much of the DNA between F56A3 and B0342 is deleted. In addition, using 6bp cutters we detect RFLPs with F56A3 and B0342 .However, the 6bp cutter RFLPs are not the same size for both cosmids. Consequently, some DNA must be present between these cosmids, which could be part of the original DNA or an insertion. Similar data as described for hDp29 and hDp24 suggest that hDp76 is extensively rearranged, deleted for K11B12 and F27E10 ,and probably does not extend to C3 OF8.
The sizes of the polymorphisms created by what we think are the endpoints suggest that ends have noncontiguous sequences added on. This is most easily illustrated with hDp29 . F56A3 detects a 173kb NotI polymorphism and a 524kb AscI polymorphism. If the duplication simply ended between F56A3 and C3 OF8,one would expect these polymorphisms to differ by 146kb (the distance between the middle NotI and AscI sites). One possible explanation for the additional DNA sequences is that the duplication ends have undergone rearrangements, for example inversions. Alternatively, the duplications could be circular or there could be DNA from elsewhere in the genome added on.
Our analysis indicates that the molecular structures of the duplications examined are extremely complex. The number and sizes of the polymorphisms do not allow us to draw a simple molecular map of the duplications, but instead suggest that they are the product of multiple rearrangements including some deletions. In addition, it appears that the duplication ends have become connected to noncontiguous DNA sequences.