Worm Breeder's Gazette 12(2): 69 (January 1, 1992)
These abstracts should not be cited in bibliographies. Material contained herein should be treated as personal communication and should be cited as such only with the consent of the author.
Exon/intron structures for both the 1.2 and 0.8 kb transcripts of the her-1 gene have been predicted from genomic and cDNA sequences (see Li et. al., this issue, for refs.). We used RNase protection assays to look for minor exons and to map the 5' and 3' ends of the transcripts present in RNA preparations from him-8 ( e1489 )animals (~40% XO). Genomic DNA fragments were cloned into the vector pT7 /T3a lpha18 and transcribed in vitro to make sense and anti-sense [32P]-labeled probes.
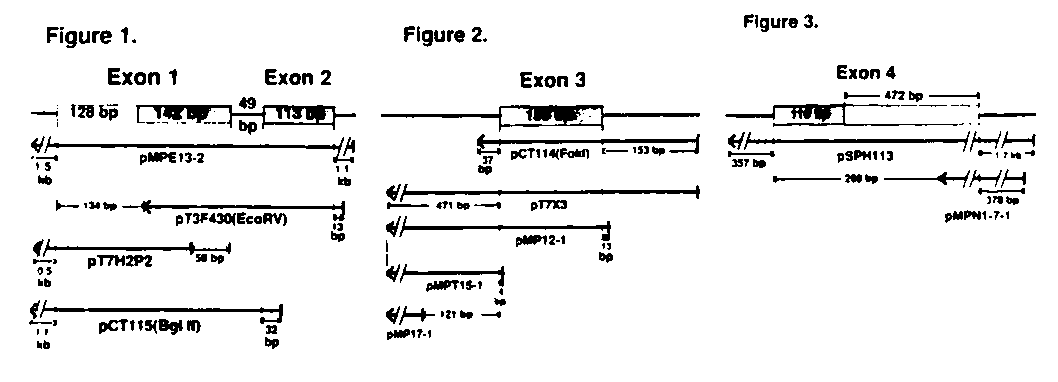
Exons 1 & 2. Probes covering exon 2 (113 bp) (pMPE13-2 and pT3 F430(EcoRV))protected one set of bands 110-114 bases long which were not protected by probes starting further upstream ( pT7 H2P2)(Fig. 1). We attribute size variations of a few bases seen within 'sets' of bands in this and other cases to the 'breathing' of the ends of probe-fragment duplexes causing variable protection of the ends. Probes covering the 5' end of exon 1 protected four sets of bands of approximately 175, 210, 235 and 255 bases (pMPE13-2 and pCT115 (BglII)),except the probe pT7 H2P2which protected fragments 55 to 70 bases smaller, suggesting multiple transcription start sites at a promoter (P1) 5' of exon 1. This was confirmed by an internal probe ( pT3 F430(EcoRV))which protected a single, much stronger set of bands of about 135 bases (Fig. 1).
Exon 3. Exon 3 (158 bp) was protected by probes from the plasmids pCT114 (FokI), pT7 X3and pMP12 -1(Fig. 2). In addition to strong bands corresponding to the full-length exon (150-165 bases), the probes pCT114 (FokI)and pT7 X3also protected a weak band of 330 bases while the probe pMP12 -1protected a weak band of 185 bases. Since exon 3 can either be spliced in cis to exon 2 or in trans to SL1 ,we reasoned that this weaker band might be the primary (unspliced) transcript of the smaller her-1 mRNA. High resolution mapping with a shorter probe (pMPT15-1) protected bands of 35 and 40 bases, but pMP17 -1,which is even more 5', did not appear to protect any bands (Fig. 2). These data suggest that intron 2 contains a strong second promoter (P2) which drives a transcription start site 2040 bp upstream of exon 3, and that enough unspliced RNA is present to be detected by the protection assay.
Exon 4. The probe pSPH113 protected bands of approximately 610, 575, 525 and 490 bases. By using a probe which started much closer to the AATAA sequences and which ended within exon 4 (pMPN1-7-1(HindIII)) we showed that all four of these fragments are from her-1 (Fig. 3). These data suggest the use of multiple poly-A addition sites in the 3' untranslated region of exon 4.
Similar experiments were done with RNA from embryos of N2 (0.2% XO) and an mnT12 strain (0.02% XO). The strong bands of exons 3 and 4 seen with the RNA from him-8 were barely detectable in RNA from either N2 or mnT12 .A single attempt to quantitate the relative amounts of signal using an AMBIS radiation detector indicated the signal from him-8 was at least 10 times stronger than that from N2 or mnT12 .The fainter bands of exons 1 and 2 and the putative unspliced exon 3 were not detected above background in N2 or mnT12 .The presence of similar amounts of the exon 3 and exon 4 transcripts in N2 and mnT12 RNAs indicates that there is a low level of transcription from the P2 promoter in XX embryos.
[See Figure 1]