Worm Breeder's Gazette 12(2): 52 (January 1, 1992)
These abstracts should not be cited in bibliographies. Material contained herein should be treated as personal communication and should be cited as such only with the consent of the author.
Genetic analysis of lin-4 has indicated that lin-4 functions at the top of the heterochronic regulatory pathway. lin-4 loss of function mutations (lf) are virtually identical to lin-14 semi-dominant gain of function (gf) mutations, causing the reiteration of early cell lineage patterns and thus the elimination of normally later cell lineage events in a number of post-embryonic lineages. Genetic and molecular analyses have shown that lin-4 is required to generate the temporal gradient of lin-14 protein, high to low from early to late in development, that determines the temporal fate of certain cells in the developing larvae. Both lin-4 (If)and lin-14 (gf)mutations cause the level of lin-14 protein to remain abnormally elevated throughout development. Gary Ruvkun's lab has mapped two lin-14 (gf)mutations to the 3' untranslated region (UTR) of the lin-14 message, suggesting a simple model where lin-4 interacts, directly or indirectly, with the 3' UTR of lin-14 to down regulate lin-14 protein levels.
In order to better understand how lin-4 functions, we have begun the molecular characterization of lin-4 .We have identified a 693 bp SalI genomic fragment that rescues the lin-4 phenotype when cloned in either orientation in the pBluescript II SK(-) vector and microinjected into lin-4 ( e912 )worms. Most of this 693 bp SalI fragment is removed by the 5 kb e912 deletion, the right end point of the deletion lies between the BglII and BstXI sites of the lin-4 SalI fragment. Further, the EMS induced lin-4 allele ( ma161 )has a C to T transition at bp 517 within the 693 bp SalI fragment. These data indicate that the sequences essential to lin-4 function are contained within this 693 bp SalI fragment.
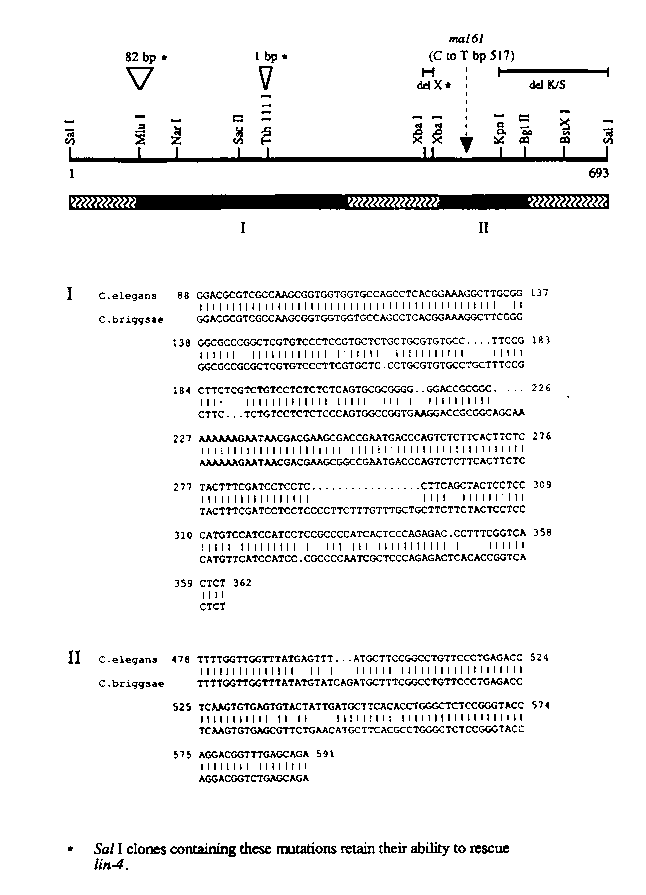
Several lines of evidence make it unlikely that the lin-4 product is a protein. Translation of the 693 bp C. elegans lin-4 DNA sequence in all six reading frames would potentially produce only small (56aa, 36aa, 12aa etc.) lin-4 proteins. It is improbable that a larger lin-4 protein is created by splicing because of the lack of appropriately positioned consensus splice sites. Further evidence against a lin-4 protein is provided by comparison of the C. elegans and C. briggsae lin-4 sequences. A 1.5 kb EcoRI genomic fragment isolated from C. briggsae is capable of rescuing the C. elegans lin-4 ( e912 )mutation by microinjection. Alignment of the C. elegans 693 bp SalI fragment with the C. briggsae 1.5 kb EcoRI fragment using the Bestfit program revealed regions of striking homology between the two species (depicted by the black boxes labelled I and II and shown in the lower half of the Figure) separated by less well conserved regions (squiggly lined boxes). However, no ORF that has both a start and stop codon is well conserved between the two species. In addition, the nature of the sequence conservation between C. elegans and C. briggsae counter indicates a protein model. Within boxes I and II there are regions of virtual sequence identity separated by gaps of one or more nucleotides rather than the third base pair substitutions that are more typical of protein coding regions. These results suggest that the lin-4 nucleotide sequence itself is critical to lin-4 function.
[See Figure 1]
Currently, we favor the idea that lin-4 encodes an RNA molecule and we are testing this by accumulating further information on the sequences required for lin-4 function. Two insertions that cause frame shifts in potential ORFs, 82 bp at the MluI site and 1 bp at the T th111 Isite, retain rescuing activity, adding support to the idea that lin-4 is not a protein. Further, a lin-4 SalI clone containing an 11 bp XbaI-XbaI deletion (del X) is also capable of rescuing the lin-4 phenotype. However, sequences to the right of the KpnI site are required for lin-4 function because a KpnI to SalI deletion (del K/S) eliminates rescuing activity. We are also attempting to determine functional lin-4 sequences by cloning lin-4 from other worm species in the hopes of identifying lin-4 sequences that have been evolutionarily conserved. Finally, we are looking for a lin-4 RNA product. Standard Northern analysis has not identified a lin-4 RNA and therefore it may be of low abundance. We are currently using oligonucleotides within lin-4 conserved regions I and II to prime cDNA synthesis followed by PCR amplification to identify transcribed sequences.