Worm Breeder's Gazette 12(1): 23 (September 1, 1991)
These abstracts should not be cited in bibliographies. Material contained herein should be treated as personal communication and should be cited as such only with the consent of the author.
The immunosuppressant Cyclosporin A is the most widely used drug to prevent graft rejection in organ transplants. However, the mode of action of cyclosporin A is not well understood. Cyclosporin A binds with high affinity to a cellular protein called cyclophilin, which has recently been shown to be identical to peptidyl-prolyl cis-trans isomerase (Fisher et al., Nature, 337, 476, 1989). Cyclophilin is highly conserved among eukaryotes, from yeast to man.
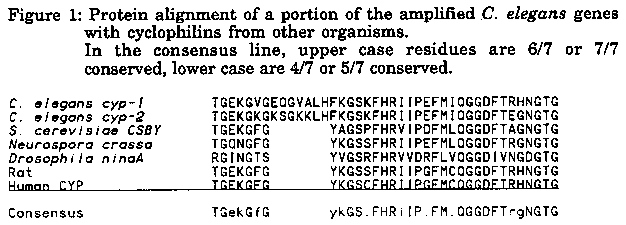
We are interested in the possibility of using C. elegans to investigate cyclophilin. As a first step, we have undertaken the cloning of worm cyclophilin genes. Using degenerate oligonucleotide primers from known conserved regions of the gene, we obtained two distinct bands on agarose gels after PCR amplification of total genomic DNA. The sequences of the amplified products revealed that these two bands represent two distinct cyclophilin genes, which we have tentatively named cyp-1 and cyp-2 (for cyclophilin). cyp-1 is 74% identical to the S. cerevisiae CSBY cyclophilin and 83% identical to the human T cell cyclosporin A-binding protein (product of the CYP gene) over the 188 amino acid region situated between the two primers, while cyp-2 shows 66% and 64% identity, respectively. The yeast CSBY and human T cell proteins are 74% identical over this region. Figure 1 shows an alignment of a portion of the amplified fragments with cyclophilins from other organisms. While the two C. elegans genes are no more homologous to each other (76% identity) than they are to cyclophilins found in other species, we think that the worm genes probably arose by duplication of an ancestral gene after the divergence of the C. elegans ancestor from the other branches, because they both contain a stretch of seven amino acids not found in other species (Figure 1). We are currently mapping these two genes onto the physical map by hybridization to the polytene YAC filters, and are screening cDNA libraries to get full-length transcripts.
[See Figure 1]