Worm Breeder's Gazette 12(1): 18 (September 1, 1991)
These abstracts should not be cited in bibliographies. Material contained herein should be treated as personal communication and should be cited as such only with the consent of the author.
We are using the polymerase chain reaction to map chromosome V deficiencies with respect to the cosmid contig map. The method was developed by R. Barstead and R. Waterston. PCR reactions are applied to deficiency homozygotes, which are arrested as either embryos or L1 larvae. Two sets of primers are used, a control set and a set from the cosmid of interest. If the deficiency deletes the site from which the second set of primers are derived, then no product will be produced. The deficiencies we are mapping have breakpoints near or to the left of unc-60 .We chose these deficiencies for two reasons. First, many of them are crossover suppressors, reducing recombination over the left half of chromosome V to less than 10% of control levels (Rosenbluth et al. Genetics 124:615). Second, we have an ongoing interest in the organization of essential genes in this region of the genome.
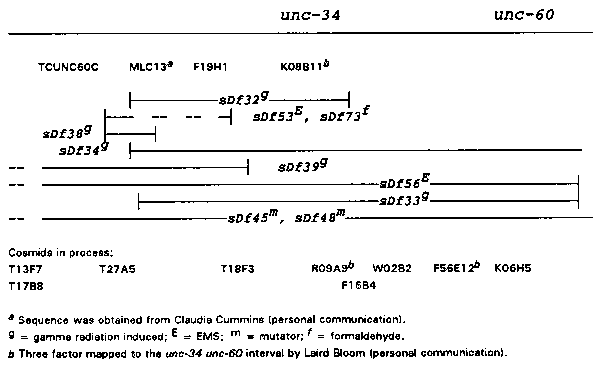
As part of this analysis, two more deficiencies have been characterized as crossover suppressors. These are sDf56 and sDf73 .The simplified map below shows the results of our mapping. We have observed some inconsistencies with the current contig map. Continued mapping will help address these problems. A simplified map is shown below; it is drawn based on the PCR data and genetic data of Johnsen and Baillie (Genetics in press). We have mapped the deficiencies with respect to four sites and are developing primers for another nine. sDf38 breaks between MLC13 and F19H1 ,a two cosmid interval. Furthermore, sDf34 and sDf39 overlap in the MLC13/ F19H1 region, yet they complement each other, indicating there are no essential genes in this region. The mapping has not shed any new light on why deletions suppress crossing over. There is no correlation between deficiency structure or the deletion of a particular site and the recombination phenotype.
[See Figure 1]