Worm Breeder's Gazette 11(5): 81
These abstracts should not be cited in bibliographies. Material contained herein should be treated as personal communication and should be cited as such only with the consent of the author.
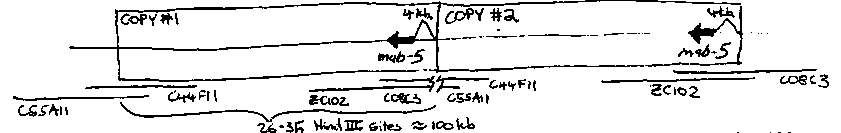
Several lines of evidence suggest that e1751 might be a gain-of- function allele of mab-5. The effect of e1751 on the Q migrations is directly opposite that of mab-5 loss-of-function mutations. (Ed Hedgecock, Dev. 100) In addition, e1751 causes generation of ectopic V ray papillae, whereas mab-5 loss-of-function alleles prevent V ray production. (Andrew Chisholm, pers. comm.) Mapping experiments by Ed Hedgecock showed that e1751 could not be easily separated from mab-5 by recombination, and finally, a restriction length polymorphism associated with e1751 was observed on Southern blots probed with the mab-5 containing cosmid ZC102. (R. Hoskins, pers. comm.) We have confirmed this observation and mapped the rearrangement: [See Figure 1] The rearrangement is a large tandem duplication of approximately 100 kb. that includes the entire mab-5 coding region and about 4 kb. of upstream sequences. As might be expected for a tandem duplication, e1751 reverts spontaneously to wild-type. These revertants no longer contain the duplication and presumable arise by recombination. We do not believe that the increased number of mab-5 gene copies explains the gain-of-function phenotype of e1751 because neither e1751/nDf16 nor e1751/mab-5(lf) is wildtype. Instead, the rearrangement may change the activity of one of the promoters. The truncation of one promoter (copy #1) 4 kb upstream of the gene could remove negative control elements or bring new enhancers close to the mab-5 gene. Some of our earlier genetic experiments suggested to us that e1751 was not an allele of mab-5. In these experiments the phenotype of e1751/nDf16 was compared to the phenotype of e1751/mab-5(e1239). Assuming that e1751 is an allele of mab-5 one would expect it to look the same over null and over deficiency. Instead, the QR migration in e1751/nDf16 is fully mutant while e1751/mab-5(e1239) has an intermediate phenotype like e1751/+. We have repeated these experiments using e2088, another allele of mab-5 that is likely to be null, and the result is the same. These results imply that something on the wild-type chromosome other than mab-5 is ameliorating the e1751 phenotype. We do not know how to interpret these results at present, but have cloned both copies of mab-5 from e1751 and plan to test their activities in transgenic lines.