Worm Breeder's Gazette 11(5): 63
These abstracts should not be cited in bibliographies. Material contained herein should be treated as personal communication and should be cited as such only with the consent of the author.
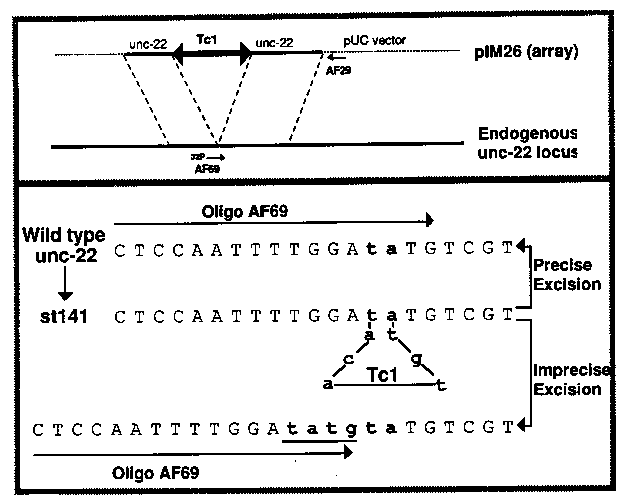
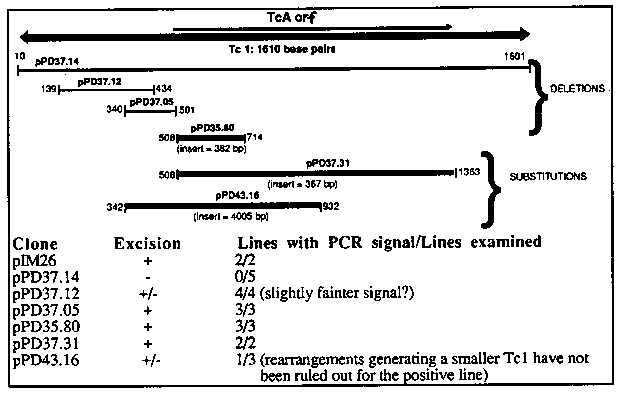
Tc1 transposition could provide a useful tool for controlled distribution of sequences around the genome. Toward this end, we have been performing Tc1 excision assays in vivo. Mori and Waterston kindly supplied a starter plasmid pIM26 (3), which has a segment of the unc-22 gene flanking an insertion of Tc1(st141). This plasmid was introduced into high copy number extrachromosomal arrays by co- transformation with the dominant selectable marker plasmid pPD10.46 ( creating 'twitcher' lines: see gazette 11#2 p20). DNA was extracted after several generations and analyzed by polymerase chain reaction ( PCR) for the presence of products indicative of excision. The transformants were in a Bristol (N2) background, so that we are presumably assaying primarily for somatic excision. We have no evidence that the Tc1 insertion in PIM26 is an autonomous element, and indeed this particular element apparently has a mutation near the 3' end of the TcA coding region, generating an extra PvuII site. Because of the possibility of rearrangements in the tandem arrays the potential for spurious PCR bands at expected mobilities, we imposed two kinds of criteria on potential PCR signals: A. A real excision signal should require both PCR primers, should be absent with pure plasmid DNA, non-transformed genomic DNA, and simple mixtures of the these. B. The spectrum of excision products should be similar to those previously observed (1,2). In particular, most Tc1 excision events leave a short residual sequence at the original insertion point. Usually this residue is the sequence TATG, derived from one end of the transposon. The two PCR primers used for the assay were as follows: an unlabeled primer (AF29) primes within plasmid vector sequences and a [32P] labelled unc-22 primer (*AF69) spanning the Tc1 insertion site. The latter oligo does not prime the parent plasmid, but should prime in cases of either precise excision or imprecise excision with the 4bp residue. Initial analysis of PCR products on agarose gels was somewhat disconcerting in that an apparent excision signals could be obtained by mixing N2 genomic DNA and the injected plasmid. This signal results from cross-annealing during PCR of extension products from the endogenous unc-22 gene and the extrachromosomal array, which mimics a homologous recombination event between the two DNAs. For this reason, we switched to sequencing gel analysis of restriction cut PCR products. At this resolution, the products of precise and imprecise excision can be separated. In vivo excision products had a distinctly different size distribution than the precise excision product artifactually generated in vitro. The bulk of the in vivo material is in a band four nucleotides larger than the precise excision product. This corresponds well with what would be expected given that in vivo excision most often leaves a 4bp residual. A fainter band is also observed in the in vivo samples at 2nt larger than the precise excision product. Possibly this reflects some excisions that leave just a 2bp residual sequence. The PCR band corresponding to imprecise excision products with a four base residual is an acceptable assay for Tc1 excision. We have constructed several internal deletions and substitutions in Tc1, introduced them into tandem arrays, and assayed for ability to excise. The derivative Tc1's and their excision abilities are shown in the table. [See Figures 1-2]