Worm Breeder's Gazette 11(5): 61
These abstracts should not be cited in bibliographies. Material contained herein should be treated as personal communication and should be cited as such only with the consent of the author.
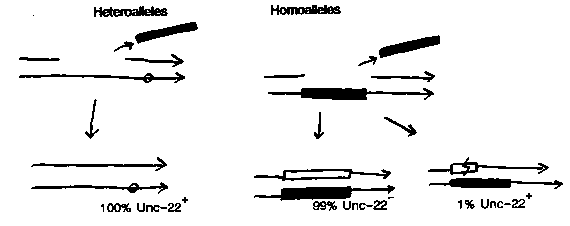
Mutations caused by Tc1 can revert, and usually a 'footprint' of a few nucleotides from one or both ends of Tc1 is left behind. This raises the question how such imprecise excision could be compatible with a reintegration of the incomplete excised Tc1 somewhere else. Inspired by recent work on P elements (Cell, 62. 515-525) that showed that efficient P element loss is homolog dependent, and by a paper by Mori, Moerman and Waterston (MGG 220 251-255) that showed that some crosses seem to activate Tc1 activity I investigated reversion of Tc1 alleles of unc-22. I found that the reversion frequency is 100x higher when one allele is a Tc1 allele, and the other allele is an EMS allele (e66). These heteroallelic strains are made by crosses (similar to those described by Mori et al.), and that - I believe- explains the apparent activation of Tc1 by some crosses. When the EMS allele is marked with a nearby visible (dpy-4 or dpy-13) the fate of both alleles in a self fertilizing hermaphrodite can be followed: I found that the homozygous EMS did not revert, the homozygous Tc1 reverts at a low frequency (10), and the heteroallele reverts at high frequency (10). This is found in four consecutive generations. The effect was found for two Tc1 alleles (st136 and st192) and for two different mutators (mut-5 and mut-6). These results suggest that repair of the double strand DNA break left after Tc1 excision depends on the presence of the wildtype sequence on the homologous chromosome. In homozygous Tc1 alleles in 99% of cases the complete Tc1 is copied back in from the other allele and no reversion is found. Only interrupted repair can -in some cases- lead to reversion. That explains the presence of a few nucleotides of Tc1 in the footprint. In other words: the excision of Tc1 is always frequent and precise, frequent reversion however depends on double strand gap repair from the homologous chromosome. A prediction is that the reversion sites from high frequency revertants in heteroallelic animals never show a footprint. I am checking that at the moment. [See Figure 1] So what is a picture of Tc1 jumping that fits all the data thus far? The first step is the precise double strand excision of Tc1. That requires trans-factors that are present in somatic cells of all worms but in the germ line only of worms that have a mutator locus. Then the excised Tc1 element goes somewhere else to reintegrate and the cell tries to repair the gap by gene conversion from the homologous chromosome. This might be described as conservative replicative transposition: conservative because the jumping element is not replicated, replicative because gene conversion copies in two new strands of Tc1 from the homologous chromosome. The net result is gain of one Tc1 element. The model can serve as a starting point for new experiments. We are reinvestigating the extrachromosomal Tc1 copies ( circles and linears have been seen by Ruan and Emmons (PNAS 81 4018- 4022): within this model they could represent intermediates in Tc1 jumping). The model may explain why our previous efforts to detect Tc1 excision in vitro were never successful: that would require efficient pairing of homologs and double strand gap repair, which is a bit much to ask in a test tube. One note: like Mori et al. I noticed a maternal effect: efficient Tc1 excision is found only when the mutator is introduced through the mother, not when it comes from the father. I think this may not have much to do with the difference in reversion rates of homoalleles and heteroalleles: it looks as if establishment of Tc1 germ line activity may take more than one or two generations after the introduction through the father of a mutator locus. We are currently studying this effect in more detail. It implies that there may be a positive feed- back in mutator expression. Note added (thanks to the FAX, Mark Edgley's divine kindness and glue): the prediction about the sequence of reversion sites was tested today. Six out of six revertants from a heteroallelic animal show precise reversion, seven out of seven revertants from a homoallelic animal show a footprint. This confirms the model presented.