Worm Breeder's Gazette 11(5): 59
These abstracts should not be cited in bibliographies. Material contained herein should be treated as personal communication and should be cited as such only with the consent of the author.
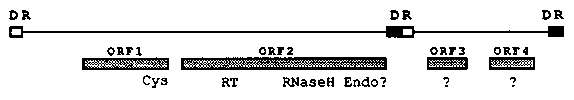
The PAT elements of Panagrellus redivivus, described earlier (de Chastonay et al., WBG 11,4), code for a major transcript (about 900nt long) which maps to the preferentially deleted portion of PAT entities. Sequence analysis of this region has revealed the presence of a single, COOH terminal cysteine motif, thought to be exclusively characteristic of retroid GAG proteins. Longer exposition of Northern blots lights up further PAT specific signals, the most noteworthy of which is an approx. 1800nt band mapping slightly downstream of the putative GAG gene. A 587 amino acid ORF, as deduced from nucleic acid studies, is found in the corresponding region. ORF2, as we refer to it here, contains a YXDD box and neighboring motifs typical for reverse transcriptase (RT). The RT region is COOH terminally followed by a tether and an RNaseH motif. Analysis of sequences further downstream suggests the presence of an endonuclease, albeit lacking a metal binding domain. No protease like motif was found in either of these ORFs. PAT ORFs 1 and 2 are on the same reading frame, but they have no overlap and the transcripts detected on Northern blot are discrete. Hence, ratio of GAG to Pol is not regulated by a translational frame- shifting mechanism but, rather, seems to be regulated at the transcriptional level. The strong transcription rate of ORF1 is paralleled by the presence of a TATA and a CAAT box, while the latter regulatory signal is not found preceding the weakly transcribed, putative Pol gene (ORF2). Two further ORFs (i.e., 3 and 4) are located further downstream, but neither one has an apparent trans- membrane domain, as one would expect from a putative Env gene of infectious retroids. These structural features put together incite us to classify PAT elements as retrotransposons, and optimal alignments with published RT sequences, as well as the order of functional domains in ORF2, seem to assign PAT to the Gypsy group of retroid elements. As described (WBG, op. Cit.), however, PAT has a split DR structure, the only precedent of this being the Toc-1 element of Chlamydomonas reinhardtii (Day et al. EMBO J. 7,1917-1927,1988). We therefore propose to dub these elements 'Para-retrotransposons', 'para' reflecting the positions of DRs if a transposition intermediate of these elements was circular. [See Figure 1]