Worm Breeder's Gazette 11(5): 55
These abstracts should not be cited in bibliographies. Material contained herein should be treated as personal communication and should be cited as such only with the consent of the author.
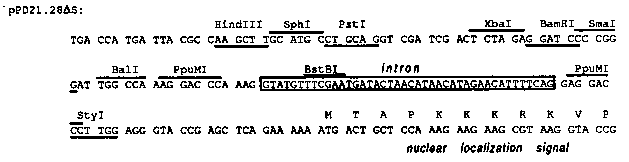
Andy Fire and colleagues (Gene 1990, Vol. 93, 189-198) developed a set of LacZ expression vectors containing a small synthetic intron, an SV40 nuclear localization signal fused to LacZ, and a 3' untranslated region derived from unc-54. For exon fusions the vectors pPD21.28, pPD22.04 and pPD22.11 provide a polylinker in all three reading frames. However, in pPD22.11 the sites upstream of the XbaI site cannot be used due to a stop codon in the XbaI site. To make some of these sites useful for cloning, pPD21.28 was digested with SalI, filled in and religated. In the resulting vector, pPD21.28deltaS, the HindIII, SphI and PstI sites can now be used in the third reading frame. [See Figure 1] In C. elegans, many exons are relatively small (~100bp), thus it happens frequently that convenient restriction sites are close to the 5' end of the exon. Fusions of these sites to the polylinker would create small exons just upstream of the synthetic intron that might not get spliced properly. To circumvent that problem, the intron cassette was deleted by digesting with PpuMI and delegating to yield the vectors pPD21.28deltaI, pPD22.04deltaI, pPD22.11deltaI and pPD21. 28deltaSdeltaI. Occasionally, convenient restriction sites cannot be found in the coding region, but are present in the intron. To make the unique BstBI site in the intron more useful for such constructs, a synthetic oligonucleotide was inserted that provides unique sites. Depending on the orientation of the oligonucleotide insertion the vectors are called LA or LB: pPD21.28LA, pPD21.28LB, pPD22.11LA, and pPD22.11LB. However, we currently do not have data that show that fusion introns get spliced properly in these vectors. [See Figure 2]