Worm Breeder's Gazette 11(5): 44
These abstracts should not be cited in bibliographies. Material contained herein should be treated as personal communication and should be cited as such only with the consent of the author.
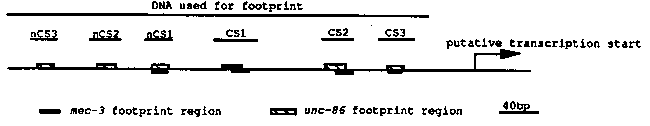
In the last newsletter, we reported that DNase I footprints showed unc-86 protein binding to three conserved sites in the upstream regions of the mec-3 gene of C. elegans and C. briggsae (CS1, CS2 and CS3). In addition gel shifts demonstrated that mec-3 protein binds to CS3. We have extended our work to search for more unc-86 binding sites using footprint analysis. In addition to the three sites previously found, unc-86 binds to three other sites (nCS1, nCS2 and nCS3). These sites are upstream of the concensus sites and are not conserved between the two species (see figure below). The mec-3 footprint using E. coli derived protein on the same region reveals four binding sites that overlap with four of the unc-86 binding sites (CS1, CS2, CS3 and nCS1, we don't know whether mec-3 binds to nCS3 and nCS2). The unc-86 and mec-3 footprint sequences are co-centered in CS3 and nCS1, but are skewed in CS2 and CS3. To test whether unc-86 and mec-3 interact with each other, we performed gel- shift analysis with CS2 and CS3 oligos. The mec-3 protein binds both oligos and shows one retarded species. In contrast, the unc-86 protein binds to both oligos with multiple retarded species, supposedly in monomer, dimer and mutimeric forms. There are three retarded species for CS2 and four species for CS3 (CS2 and CS3 do not share sequence homology; CS2 has POU binding motifs, while CS3 is AT- rich). When both proteins are added together in the presence of excess probe, the gel-shift pattern does not change with the CS2 oligo. However, the CS3 gel-shift pattern is changed. Though the mec-3 band, which has faster mobility than all unc-86 bands, and the second fastest unc-86 band appear unchanged, the remaining unc-86 bands have disappeared. A new band appears at a position between the two fastest original unc-86 bands. These results suggest that the interaction of mec-3 and unc-86 proteins inhibits the formation of the multimeric complexes by unc-86 protein. Also the loss of the fastest unc-86 species (supposedly the monomeric form) and the appearance of a new band at a slightly higher position (supposedly a heterodimeric form) by the addition of mec-3 protein in the presence of excess probe suggest that the binding of the unc-86 and mec-3 monomer to CS3 may be cooperative. We don't know the in vivo significance of this phenomenon, but are testing its importance by in vitro mutagenesis. We are also using in vitro mutagenesis to knock out the unc-86 and mec-3 binding sites. Changing five nucleotides in CS3 completely abolishes the unc-86 and mec-3 binding to this region as verified by unc-86 footprinting and mec-3 gel-shift. When this mutant is introduced into wild-type animals as a mec-3-lacZ construct, the staining is almost completely abolished, suggesting that CS3 is essential for mec-3 expression. Three nucleotide changes in CS1 abolish the unc-86 binding at this region (these changes are not in the mec-3 footprint region). This mutant seems to have no effect on staining in ALMUR and PLMUR, but greatly reduces the frequency of staining in FLPL/R, and PVDUR (we are not quite sure about AVM/PVM), suggesting that this CS is important for the expression of mec-3 in FLP and PVD cells (non-touch-cell). Preliminary results of similar CS2 mutants suggest that they affect ALM and PLM staining but not PVD and FLP (ALM staining is abolished; PLM staining is greatly reduced). Thus, CS1 and CS2 may be important for cell-specific mec-3 expression. Deletion of a region which contain the three non-conserved unc-86 binding sites leads to the appearance of additional staining cells, suggesting that this region contains negative control elements. Perhaps unc-86 binding (or mec-3 binding ) at this region could repress mec-3 expression in these other cells. Moreover, these results suggest a caution in relying too heavily on conservation as the sole method of identifying important cis acting elements. [See Figure 1]