Worm Breeder's Gazette 11(5): 36
These abstracts should not be cited in bibliographies. Material contained herein should be treated as personal communication and should be cited as such only with the consent of the author.
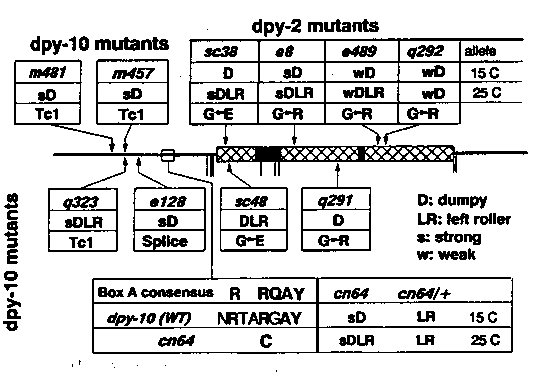
dpy-2 and dpy-10 are tightly linked at the left end of the cluster of LGII. Alleles of each display genetic interactions with other loci. The dpy-2 alleles e8 and q292 and the dpy-10 alleles e128 and q291 can suppress a temperature sensitive allele of glp-1 (Maine and Kimble, Development 105: 133). e128 can suppress a ts allele of mup-1 (T. Bogaert, personal communication). Kusch and Edgar (Genetics 113: 621) showed interactions between the recessive Dpy allele dpy-10(e128) and the recessive Lrol allele sqt-1(sc13): e128 sc13/e128 + animals are DpyLRol. We have since shown that at least three other sqt-1 alleies ( sc99, sc101, and sc1) behave in a similar manner in the e128 background such that e128 sqt-1/e128 + animals are DpyLRol. e128 sc13 homozygous animals are DpyLRol. All other e128 sqt-1 homozygotes tested were Dpy, however the character of the Dpy phenotype differs with different sqt-1 alleles. Cosmid ZK857 is located in the predicted vicinity of dpy-10 and contains two collagen genes. Using mutator induced alleles of dpy-10 ( kindly provided by Don Riddle and Tim Schedl), we were able to show that one of these collagens is dpy-10. We have shown that the other is dpy-2 by sequencing of mutants. dpy-2 and dpy-10 are located 3.5 kb apart (dpy-2 on the left) and are transcribed in opposite directions. Structurally, the dpy-2 and dpy-10 collagens are more similar to each other than to any other of the sequenced collagens and are probably a result of a gene duplication. dpy-2 is trans-spliced with SL1; dpy-10 is not trans-spliced. The point and Tc1 mutations that we have analyzed are diagrammed in the figure below. Glycine Substitutions: Replacements of Gly with Arg or Glu in the Gly-X-Y repeat region were identified in alleles of both dpy-2 and dpy- 10. Different substitutions produce different phenotypes and some are temperature sensitive. In general, temperature affects the severity of the LRol phenotype much more than the Dpy phenotype. The locations of the substitutions apparently dictate the Dpy or DpyLRol phenotype. These substitutions presumably result in the destabilization of the collagen triple helix, which requires a glycine at every third amino acid for proper folding. A Splice Acceptor Mutant: dpy-10(e128) is a G->A transition in the conserved AG of the splice acceptor of intron 2. Reverse transcription PCR of e128 RNA indicates the presence of approximately equal amounts of apparently normally spliced message and unspliced message. Even though the AG of the splice acceptor sequence is normally required for splicing, the mutant acceptor appears to be utilized surprisingly well. When intron 2 is not spliced a 16 amino acid insertion would be encoded by the intron (no nonsense codons or frameshift). Whether the Dpy phenotype of e128 is a result of reduced level of normal dpy-10 product or the production of dpy-10 collagen with a 16 amino acid insertion is unknown. No evidence for the utilization of cryptic splice acceptors was detected. Another Dominant Gain of Cysteine Mutant: dpy-10(cn64) displays dominant LRol and recessive temperature-sensitive DpyLRol phenotypes. The cn64 mutation results in an Arg->Cys substitution in homology box A, the same Arg->Cys change seen in the dominant RRol sqt-1(e1350) and rol-6(su1006) mutations. All dominant Rol mutations identified to date are Arg->Cys changes in the A Box. However, the dpy-10 Arg->Cys mutation causes a dominant LRol phenotype while the sqt-1 and rol-6 mutations are dominant RRol. Thus, the equivalent mutation in different collagens can cause twisting in opposite directions. These gain of Cys mutations probably allow the formation of abnormal disulfide bonds within the cuticle. Tc1 Insertions at the Same Site Produce Different Phenotypes: Two of the three Tc1 insertions in dpy-10(q323 and m457) are located at exactly the same point in the gene, and are in the same orientation. It is interesting that these two strains have different phenotypes, q323 is DpyLRol and m457 is Dpy. One possible explanation for the difference in phenotypes is that the excision rates of the two elements are different. We examined the relative frequency of excision in the three Tc1 strains by PCR of individual worms. We found that the amount of the excision-sized product is consistently about ten-fold less with q323 than with m457 or m481, indicating that the element in q323 excises less frequently. The amounts of excision in m457 and m481 are approximately equal. The lower frequency of excision in q323 animals may result in the production of less functional dpy-10 product than in m457 or m481 animals, resulting in the DpyLRol phenotype. It is possible that the differences in excision rates may be caused by differences in the sequences of the elements. However, the effect could also be due to dpy-10 linked differences In the genetic backgrounds of the strains. ? Null phenotypes: Our observations suggest that the null phenotypes of dpy-2 and dpy-10 are DpyLRol and that the less severe phenotype is Dpy. Analysis of Tc1 insertions indicate that a lower amount of excision Is correlated with the DpyLRol phenotype. In addition, the temperature sensitive alleles of dpy-2 and dpy-10 show DpyLRol phenotypes at 25 C, but are Dpy nonLRol at 15 C. Raising the temperature presumably causes destabilization of the triple helix, reducing the level of functional dpy-2 or dpy-10 collagen. Since neither dpy-2 or dpy-10 are haploinsufficient, 50% of the normal level of these collagens is presumably sufficient for normal function. The Dpy phenotype would appear when the level of functional gene product was some unknown amount less than 50%, whereas the DpyLRol phenotype would appear to represent a more severe reduction or compete absence of these collagens. [See Figure 1]