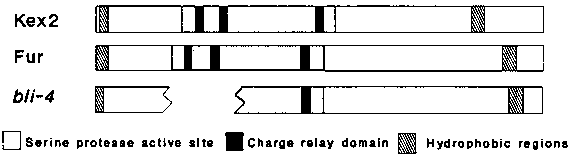Worm Breeder's Gazette 11(5): 28
These abstracts should not be cited in bibliographies. Material contained herein should be treated as personal communication and should be cited as such only with the consent of the author.
bli-4 was cloned by Tc1 tagging (WBG 11 #2). We probed staged specific and total RNA with a bli-4 cDNA. In total RNA two bands are detected: 2.7 and 4.5 kb. bli-4 is expressed throughout the life cycle, but most highly in L1 and L4/adult worms, and least in L2. We have partially sequenced cDNAs that includes all of the coding region. Searching the SWISSPROT protein database revealed similarity to the yeast gene Kex2 as well as two human genes, Fur and PC2, and a K. lactis gene Kex1. Kex2 encodes a 814 amino acid membrane bound serine endoprotease that activates by cleavage the alpha-mating factor and the killer toxin pro-proteins in the golgi body prior to secretion of these proteins (reviewed by R. Fuller A. Brake and J. Thorner, Microbiology 273-278, 1986). No function for PC2 or Fur has been determined. We have sequence corresponding to 701 amino acids of bli- 4, including the amino and carboxy ends. About 300 bp in a region representing about half of the Kex2 active site remains to be sequenced, shown as a gap in the figure below. In 70 amino acids around the sequenced portion of the active site, bli-4 is 51% similar to the Kex2 gene, 55% similar to PC2, and 68% similar to Fur.Outside of the active site the sequence identity is not strong. However, a number of structural features are conserved in all members of the Kex2 gene family, (illustrated below). These include a hydrophobic sequence at the amino terminal that functions as a leader sequence in Kex2 and a hydrophobic domain at the carboxy end that functions in Kex2 as a transmembrane domain (the latter is not present in the PC2 sequence). In addition, the spacing of these domains is conserved. We probed EcoRI digested genomic DNA from seven alleles of bli-4. Of these seven, two alleles have rearrangements. The Tc1 allele h1010 contains a Tc1 element inserted in or just 3' to the active site. e937 has a 3 kb deletion fusing two adjacent EcoRI fragments at the 3' end of the gene. We do not know if any of the coding region is deleted. e937 is hypomorphic, and is only viable allele: all of the other alleles are early larval lethals. These results suggest that the role of bli-4 in the cuticle may involve the processing of some cuticle components or some regulatory signal required for cuticle secretion. [See Figure 1]