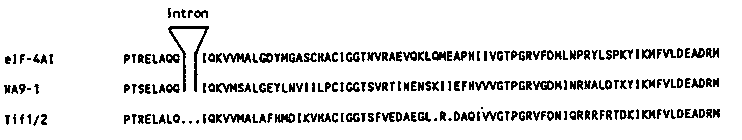Worm Breeder's Gazette 11(5): 26
These abstracts should not be cited in bibliographies. Material contained herein should be treated as personal communication and should be cited as such only with the consent of the author.
With one set of primers designed to amplify the Drosophila vasa- equivalent from C. elegans, we produced several distinct PCR products. Because these primer sequences were not unique to the vasa sequence but rather were shared by all members of the so-called DEAD family of helicases, our results were not unexpected. We predicted that the different PCR products would correspond to different helicase protein genes. The multiple PCR amplified DNA fragments were gel isolated, cloned and sequenced. To date, sequence analyses of our PCR clones suggest that C. elegans may have as many as six different potential helicase protein genes that belong to the DEAD family of helicases ( Nature 335:611-617). Most of these represent potential helicases that have not been previously characterized in another organism. It may be possible to characterize a whole family of putative helicase genes in Caenorhabditis.One of our PCR clones, called NA9-1, has 61% perfect amino acid homology to eIF-4A, the protein synthesis initiation factor that has been cloned from mouse (Nielsen et al., NAR 13:6867-6880). The identity is 83% with conserved amino acid changes. Shown below is a comparison of NA9-1 predicted amino acid sequence with that of the mouse genes eIF-4AI/II and the yeast equivalents Tif1/2 (Linder and Slonimski, NAR 16:10359). There is 54% perfect amino acid homology between the C. elegans clone, NA9-1, and the yeast Tif1/2. The position of an intron whose exact location is conserved between the mouse and nematode sequences is also indicated. The eIF-4A protein is well characterized (for review see Moldave,K., Ann. Rev. Biochem. 54:1109-1149). It recognizes the 5' cap structure of RNAs, has RNA binding activity, and has an ATPase- dependent helicase activity upon RNAs. Some of the different helicase motifs that are conserved among this large class of proteins have been functionally analyzed with eIF-4A. The two genes eIF-4AI and II, although highly conserved, have different patterns and levels of expression in different tissues (Nielsen and Trachsel, EMBO J. 7:2097- 2105). We are not yet certain if there are multiple Caenorhabditis eIF-4A-like genes. There are two EcoRI fragments of C. elegans DNA, 2.4 Kb and 1.3 Kb, that hybridize to the insert of NA9-1 on genomic Southerns, but there is also an EcoRI site in our PCR clone. In addition, the small size and unequal hybridization signals of the two genomic EcoRI fragments does not distinguish between a single gene or multiple highly conserved genes that corresponds to our PCR clone, NA9- 1. We have also used the NA9-1 insert DNA to isolate a cDNA from a C. elegans cDNA library previously constructed in this laboratory. We are currently completing the sequence analysis of this clone. Preliminary data show that the cDNA clone, called CEB1, corresponds to the amplified PCR product and that its homology at the amino acid level with eIF-4A extends outside the region delimited by the PCR primers. [See Figure 1]