Worm Breeder's Gazette 11(5): 22
These abstracts should not be cited in bibliographies. Material contained herein should be treated as personal communication and should be cited as such only with the consent of the author.
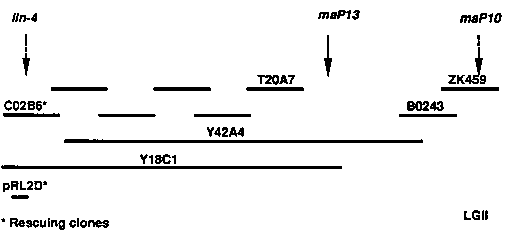
We genetically mapped the lin-4 locus to a position closely linked and to the left of bristol/bergerac polymorphism maP13, detected by YAC Y42A4. Using YAC and cosmid clones in the region as probes to southern blots, we analyzed DNA from the only lin-4 allele available at the time, e912, (see Feinbaum, et, al, this WBG issue) and found that the YAC Y18C1 detected a polymorphism in a 5 kb EcoRI band. Cosmids drawn under Y18C1 on the physical map did not detect this band, so we suspected that lin-4 was at the right end of Y18C1, which extended into a cosmid-free gap. Therefore, to clone the sequences corresponding to the 5 kb EcoRI band, we used Y18C1, which detects the 5 kb RI band, and Y42A4, which almost completely overlaps Y18C1 but does not detect the band, in a differential screen of an EcoRI lambda library prepared by Ann Rougvie. In this way we isolated two different 5 kb EcoRI fragments that both turn out to be partially deleted in lin-4(e912) DNA. Much to our surprise, these clones lie not in the gap, but in the contig and to the left of the previous end of Y18C1. We have therefore revised the extent of Y18C1 to include the cosmids containing these two 5 kb EcoRI fragments. One of the polymorphic 5 kb EcoRI clones, pRL2D, rescues the mutant phenotype of lin-4(e912). (Two overlapping cosmids, C02B6 and C31G11, also rescue lin-4). Therefore, pRL2D contains a functional lin-4 gene, although the lin-4(e912) lesion, originally generated by [32P] decay, is somewhat complex, involving deletion of another 5 kb EcoRI fragment in the region, and partial duplication of a 3.5 kb fragment. We are currently sequencing pRL2D and testing smaller subclones for rescue.