Worm Breeder's Gazette 11(5): 21
These abstracts should not be cited in bibliographies. Material contained herein should be treated as personal communication and should be cited as such only with the consent of the author.
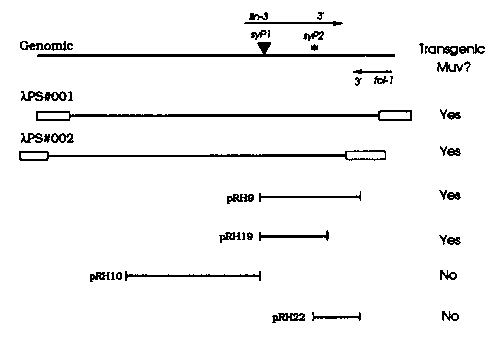
lin-3 is required for vulval induction; therefore lin-3 is either necessary for the generation of the inductive signal by the anchor cell or for the response to the inductive signal by the VPCs. lin-3 is interesting to us because genetic experiments indicate that lin-3 acts early in the pathway of vulval induction and because lin-3 appears to be essential for vulval induction. Other lin-3 phenotypes include larval lethality, hermaphrodite sterility, and a Mab defect that includes crumpled spicules. The physical location of the lin-3 locus was identified by transposon tagging as described previously (WBG 11.3). We have defined a region sufficient for rescue of lin-3 by microinjecting genomic subclones in the vicinity of two lin-3 RFLPs, syP1 and syP2. lambda PS#2 rescues the lethal phenotype of the lin-3 lethal alleles n1059, s751, sy51, sy52, and sy53. lambda PS1, lambda PS2, and pRH9 can all rescue the vulvaless phenotype of lin-3. Surprisingly these transgenes also cause a variable fraction of lin-3(+), and lin-3(null) animals to a have a multivulva (Muv) phenotype. The multivulva phenotype could be caused by over-expression of wildtype lin-3 or some mutant effect of the transgenic lin-3 DNA. pRH10 and pRH22 have no ability to rescue and do not cause a multivulva phenotype. pRH19, which is a truncated derivative of pRH9, rescues less well than pRH9. pRH19 was used to isolate a lin-3 cDNA because it does not overlap the poxy and abundant fol-1 (friend of lin-3) transcript (which was previously detected by reverse northern analysis and shown to reside in a region not necessary for rescue of lin-3). The lin-3 cDNA was isolated from a library made by Chris Martin. This cDNA detects both lin-3 RFLPs on southern blots and is single copy in the genome. The 3' end of this cDNA extends outside of pRH19 but is within pRH9. We conclude that pRH9 contains the lin-3 coding region and that pRH19 is a crippled 3' deletion of the gene. We are sequencing lin-3 and examining the genetic properties of the transgenic lin-3 Muvs.[See Figure 1]