Worm Breeder's Gazette 11(4): 8
These abstracts should not be cited in bibliographies. Material contained herein should be treated as personal communication and should be cited as such only with the consent of the author.
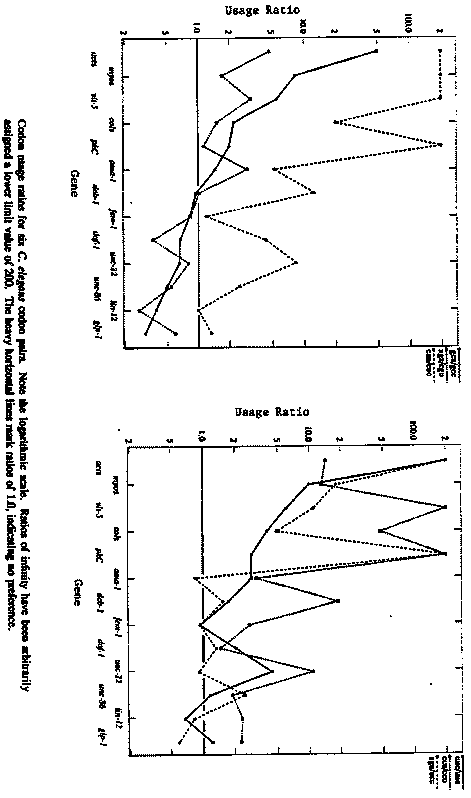
A general trend from highly asymmetric codon usage in strongly expressed genes to less asymmetric codon usage in weakly expressed genes is observed in many organisms (P. Sharp et al., Nucl. Acids Res. 17 (1988) 8207-8211). Sequence data are now available for worm genes representing a number of gene families, with a variety of expression patterns; hence one can look for variations from the distinctively biased 'average' C. elegans codon usage pattern (WBG Jan. 1990). Large differences in codon usage are not apparent between members of the myosin, collagen, actin, and lin-12-glp-1 gene families; however, there are some striking differences between gene families. Some of these are summarized in the figures on the following page, which show usage ratios for pairs of Ala, Arg, Leu, Phe, Pro, and Ser codons. In every case, the usage asymmetry is generally higher in the actin, myosin, and collagen families, and lower in unc-86 and the lin-12-glp- 1 family. The usage ratios for the Ala and Arg pairs shown (gca/gcc and aga/cgu) actually reverse, indicating a shift in codon preference. The most significant shift is in the Arg codons of daf-1,lin-12, and glp-1, in which terminal purines are preferred over terminal pyrimidines. The preference for terminal a's in Gly and Pro codons, however, is decreased in these genes relative to that found in the actins and collagens, so the shift is not a general one towards third- position purines. These data - especially the strong reversals in preference - suggest that codon usage asymmetry, at least for some amino acids, may play a role in regulating gene expression. They also make it fairly clear that codon usage asymmetry will be a poor guide to the identification of coding exons in some genes. It will be interesting to see whether other ways of looking at codon data reveal additional patterns. Sources of the data are as follows: act-1, act-2, act-3, act-4 (Krause etet a al., J. Mol. Biol. 208 (1989) 381-392); ama-1 (Bird and Riddle, Mol. Cell. Biol. 9 (1989) 4119-4130); col-1, col-2 (Kramer et al., Cell 30 (1982) 599-606); col-6, col-7, col-8, col-14, col-19 (Cox et al., Gene 76 (1989) 331-344); daf-1 (Georgi et al., Cell 61 (1990) 635- 645); deb-1 (Barstead and Waterston, J. Biol. Chem. 264 (1989) 10177-10185); dpy-13 (von Mende et al., Cell 55 (1988) 567-576); fem-1 ( Spence et al., Cell 60 (1990) 981-990); glp-1 (Yochem and Greenwald, Cell 58 (1989) 553-563); lin-12 (Yochem et al., Nature 335 (1988) 547- 550); myo-1, myo-3 (Dibb et al., J. Mol. Biol. 205 (1989) 603-613); pkC (Gross et al., J. Biol. Chem. 265 (1990) 6896-6907); sqt-1 (Kramer et al., Cell 55 (1988) 555-565); unc-15 ( Kagawa et al., J. Mol. Biol. 207 (1989) 311-333); unc-22 (Benian et al., Nature 342 (1989) 45-50 and this WBG); unc-54 (Karn et al., Proc. Natl. Acad. Sci. USA 80 (1983) 42534257); unc-86 (Finney et al., Cell 55 (1988) 757-769); and vit-5 (Spieth et al., Nucl. Acids Res. 13 (1985) 7129-7138). [See Figure 1]