Worm Breeder's Gazette 11(4): 46
These abstracts should not be cited in bibliographies. Material contained herein should be treated as personal communication and should be cited as such only with the consent of the author.
N2 carries a tandem duplication of much of the unc-30 gene; the
duplication is absent in B0. Sequencing of subclones of the N2
genomic lambda clone CB#RH17 reveals a first copy 5,592 bp in length,
followed directly by 3,387 bp of a perfectly repeated second copy. (
This is potentially dangerous, as there is the slight possibility of
gene conversion in the lambda clone.) At this position (a Sau3A site),
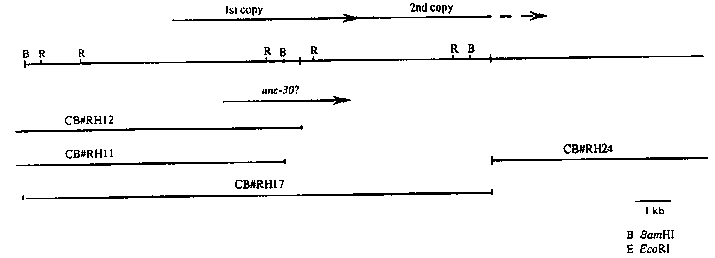
CB#RH17 ends and another genomic lambda clone, CB#RH24, begins (see
figure). Partial sequencing of subclones of CB#RH24 indicates that
the second copy continues, probably to completion. (Sequence obtained
so far includes a fragment which presumably represents the end of the
second copy; it matches the end of the first copy and then diverges.)
Much and perhaps all of the coding potential of the gene is duplicated.
CB#RH12 and CB#RH17, both of which complement unc-30(e191) in
transformation rescue experiments, end at Sau3A sites at equivalent
positions in the first and second copies, respectively. I have
previously reported that CB#RH12 breaks within an intron near the 3'
end of the gene, and thus apparently directs synthesis of an altered
but still functional product. In CB#RH17, the second copy might also
direct synthesis of such a product, unless required 5' sequences are
missing. (The ends of the unc-30 transcript have not been defined.)
Since CB#RH12 contains none of the second copy, it seems probable that
the first copy represents the real unc-30 gene, but this is not
formally proven. Further comment awaits completion of the sequence of
the second copy, which may reveal differences and an assay which
distinguishes the two copies. This may all be irrelevant to the
prediction of the amino acid sequence of the putative unc-30 protein.
By genetic criteria, only one of the two copies of unc-30 seems to
function. The six ems-induced alleles reported by Brenner ('74)
appeared to arise at about knock-out frequency. The eleven mutant
alleles of unc-30 of which I am aware are all simple recessives. I
have constructed all the heteroallelic combinations of ten alleles,
and no pair complements. (nT1 [unc-30(e2327)] wasn't tested. Two
alleles, e2384 and bx7, were induced on B0 chromosomes, which contain
only one copy of the gene.)
The first 3.4 kb of the duplication, and probably more, are
perfectly repeated, indicating that it arose recently in C. elegans
evolution. I have examined a set of wild strains by Southern blotting.
The Anglo-American strains N2 (Bristol), DH424 (California) and
TR403 (Wisconsin) carry the duplication; the Continental strains B0 (
Bergerac) and RC301 (Freibourg) carry a single copy. (Does anyone
have a strain from Martinique, or Louisiana! ?)
Given the duplication, one might expect that certain unc-30
mutations would revert by gene conversion. A strain carrying the
allele e596 appears to have reverted during passaging, but since I
have no molecular assay for the allele, I cannot exclude the
possibility of contamination by an N2 animal. The 'revertant' is
fully wild-type, retains the duplication, and is tightly linked to unc-
30. Incidentally, e596 is incompletely, but significantly suppressed
by smg-1(e1228).The position of the unc-30 transcript in the figure is
based on analysis of the sequences of CB#RH17 and two cDNA clones.
The predicted stop codon is 503 bp upstream of the beginning of the
second repeat.
[See Figure 1]