Worm Breeder's Gazette 11(4): 29
These abstracts should not be cited in bibliographies. Material contained herein should be treated as personal communication and should be cited as such only with the consent of the author.
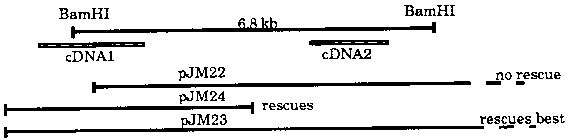
We have rescued the lin-15 multivulva.(Muv; visible or AB) and synthetic (syn; class A and class B) Muv phenotypes with cosmid PS#74B3. This cosmid maps to the far left (formerly, far right) end of the clb-1 contig on the X chromosome and was originally isolated because it weakly hybridizes to the G-protein PCR clone 1, as reported in WBG11(2):32. As we had been trying to clone lin-15 (see WBG11(3) :60 for last update), we knew from the map data that lin-15 should lie either at the right edge of the clb-1 contig or in the gap off the end. Since PS#74B3 appeared to map closely to lin-15 on the physical map and because the putative G-protein was a candidate gene for a genetic locus presumably involved in signal transduction, we decided to microinject the cosmid to test whether it rescued lin-15. As we had hoped, PS#74B3 rescued lin-15, although the G-protein hybridizing region did not rescue. A 'reverse northern' analysis identified a 6.8 kb BamHI fragment which lay within the region defined as necessary for lin-15 function by injection. When the 6.8 kb fragment was used to screen Stuart Kim and Chris Martin's cDNA libraries, we identified two classes of non-cross hybridizing cDNAs which mapped to opposite sides of our 6.8 kb fragment, with cDNA2 apparently more abundant than cDNA1. [See Figure 1] Subsequent injection results showed that genomic DNA containing cDNA1 but not cDNA2 was sufficient to rescue the Muv phenotype. Also, RFLP analysis picked up a small deletion in the lin-15(n767) class A allele in the region of cDNA1. DNA which does not contain cDNA1 but does contain cDNA2 did not rescue the lin-15(n765) Muv phenotype (n765 is Muv at >20 C, class B at 15 C)--however, an RFLP was found in n765 to the right of cDNA2. Also, genomic DNA which contains both cDNA1 and cDNA2 rescues the Muv phenotype much better than DNA containing only cDNA1. Furthermore, all of the four EMS induced visible Muv alleles plus the three lin-15 alleles generated by Stuart Kim in the TR679 mutator strain contain large deletions of DNA in this genomic region, with n377 deleting the 6.8 kb region entirely. We believe that the lin-15 locus contains at least two independently mutable regions such that a hit in one causes a class A syn Muv phenotype while a hit in the other causes a class B syn Muv phenotype, the 'two hit model'. Single hits in both regions or a large deletion covering both regions are necessary for a visible Muv phenotype. This model is supported by the facts that: 1. the four EMS induced visible Muvs contain large deletions even though the frequency of EMS induced deletions is believed to be no greater than 10%, suggesting a strong selection for deletions in lin-15 visible Muv hunts. 2. we were unable to recover a null allele in a non-complementation screen of 35, 000 F1 worms using EMS (see WBG11(3):60 for details), suggesting that single point hits can not cause a null. 3. DNA covering the n765 polymorphism is not sufficient to rescue the Muv phenotype, suggesting there may be a second hit elsewhere in the region. 4. although it doesn't seem possible to obtain visible Muv mutations as EMS single point hits, visible lin-15 alleles are easily obtained in F2 screens for syn Muvs starting with an opposite class syn Muv allele. A visible lin-15 allele recovered in a syn Muv screen using n767 only shows the deletion originally found in n767, suggesting there is a second point hit elsewhere in the region. It is possible that cDNA1 and cDNA2 are differentially spliced 3' exons of a common 5' end, or that cDNA1 and cDNA2 are driven by the same control region. Another possibility is that the lin-15 locus could comprise two closely linked genes, one which is a class A syn Muv and the other which is a class B syn Muv. We are working to resolve these issues and also to further elucidate how lin-15 functions in the vulval development pathway by further characterization of the cDNA clones and the associated genomic DNA surrounding those regions.