Worm Breeder's Gazette 11(4): 115
These abstracts should not be cited in bibliographies. Material contained herein should be treated as personal communication and should be cited as such only with the consent of the author.
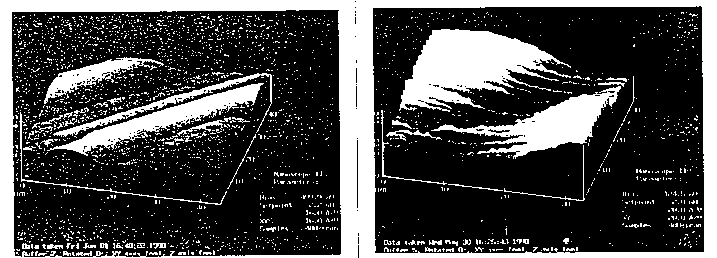
We have been studying C. elegans microtubules from cellular and genetic aspects. Since scanning tunneling microscopy (STM) can provide atomic resolution of metals and semiconductive surfaces; our objective is to obtain images of C. elegans microtubules resolved at the atomic level. However, application of STM to biomolecules is faced with a variety of problems, including poor electrical conductivity, flexible stability under STM imaging conditions. In spite of limitations, recently, impressive images of DNA biomolecules have been obtained at atomic resolution. Microtubules (MT) were isolated from 30g of freshly grown N2 worms according to the procedure of Aamdot and Culotti 1986, and Siddiqui et al 1989, using cycles of temperature dependent assembly and disassembly (Shelanski, et al. 1973). Assembled MT were taken in MT isolation buffer (containing 0.1mM GTP, 20 M taxol, and 2.0 mM DTE), diluted tenfold and fixed in glutaraldehyde (0.1%). The samples were kept frozen at -20 C, for 4-5 days. The STM imaging samples were thawed, and a drop was smeared onto a chip of freshly cleaved HOPG ( highly oriented pyrolitic graphite, Union Carbide), and allowed to air dry at room temperature. The STM images were obtained using 'Nanascope II Digital Instruments, California (Sam Howell and C. Lee, kindly helped with the STM settings). All STM imaging was conducted at atmospheric pressure and at room temperature (22 C). Two different STM images of C. elegans MT are shown here. [See Figure 1] Most of our STM scans of MT correspond to one or two microtubules, but they appear flattened and bent along the seams of the protofilaments. The dimensions of alpha and -tubulin dimer subunits are about 8 nm, and each protofilament has a width of 4.6 nm. The flattening and buckeling of protofilaments along the seams could arise due to the fixation conditions, or caused by the STM current flow through the specimen. At present, we are unable to attribute this observation to any single factor, and the images of MT do not provide any higher resolution than conventional electron microscopy using negative staining. But it is increasingly possible that one may be able to resolve individual amino acids or even atoms using STM. We plan to continue looking for improved conditions for better STM or AFM images of C. elegans MT. Brief description of STM: Scanning tunneling microscopy (STM) traces three dimensional images of surfaces - even at the level of individual atoms. Gerd Binnig & Heinrich Rohrer of IBM Zurich received the Nobel Prize in 1986 for STM (Refer, e.g.: Scientific American, August,1985). Briefly in STM, the 'aperture' is a small tungston probe, its tip extremely fine (about 0. 2 nm in width, and may consist of a single atom). Piezoelectric controls move the tip within a nanometer or two of the surface of the conducting sample, allowing an overlap between the electron clouds of the atom at the probe tip and the closest atom of the sample placed on the surface. As a small voltage is applied to the tip, electrons 'tunnel' across this gap, creating a tiny tunneling current. The strength of the tunneling current is exponentially sensitive to the width of the gap (about an order of magnitude per angstrom). Depending on the substrate, typical currents and voltages are in the range of nanoamperes and millivolts. A servo system uses a feedback control that keeps the tip to substrate gap constant, by modulating the voltage across a piezoelectric positioning system (Piezoceramic materials expand or shrink extremely small distances (i.e. angstroms ) in response to the applied voltage. As the tip scans the surface, variations in this voltage, when plotted correspond to surface structure. I wish to thank members of Ward lab (Alicia, Andrea, Bill, Bonnie, Bruce, Jacob, John and Paul); Hameroff Lab (Dolly, Larry, Mohammad, and Richard); and Sarid Lab (Sam and Lee).