Worm Breeder's Gazette 11(4): 10
These abstracts should not be cited in bibliographies. Material contained herein should be treated as personal communication and should be cited as such only with the consent of the author.
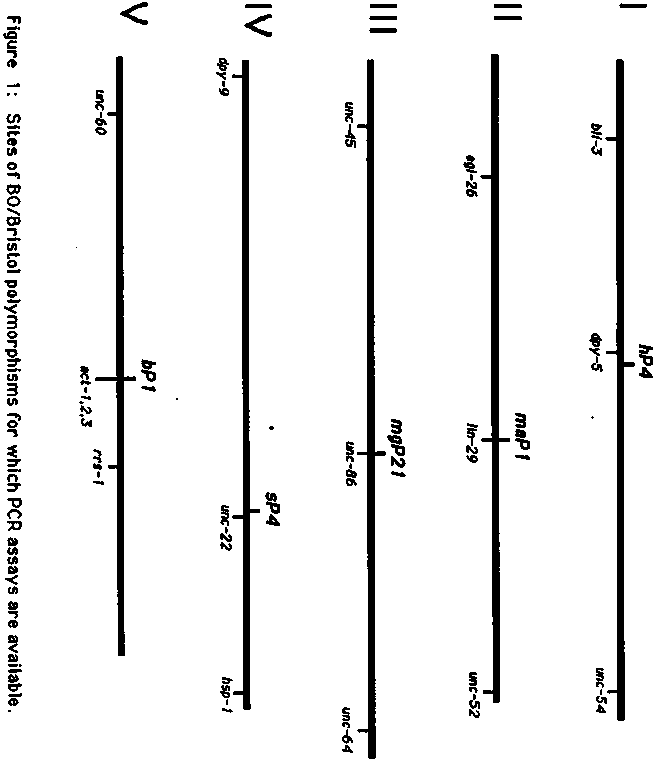
After mapping many lethal mutations by standard methods, we decided to try another approach to genetic mapping which has worked well and may be of general interest. Polymerase chain reaction (PCR) assays are being developed to specifically detect several Bristol/Bergerac Tc1 polymorphisms on each of the 5 autosomes (Fig.1). The segregation of these polymorphisms can be followed in the progeny of Bristol/Bergerac hybrids by doing PCR reactions with DNA from single worms. Each assay consists of a unique primer flanking the Tc1 insertion site and a common primer complementary to internal Tc1 sequence. The Bergerac allele is detected by amplification of a DNA fragment of characteristic size; no amplification occurs from the Bristol allele. For more efficient mapping, flanking primers were chosen so that amplification products from different polymorphisms are easily distinguishable by size on a non-denaturing acrylamide minigel. This allows several assays to be used in the same PCR reaction. To map a mutation, heterozygous Bristol males are crossed into the Bergerac strain RW7000, and an F1 hermaphrodite that segregates mutants is isolated. 20 - 30 of F2 mutant homozygotes are picked (dead or alive!) to separate tubes and their DNA is subjected to PCR analysis with the assay set of primers (see below). For unlinked polymorphisms, the Bergerac allele is detected in 75% of the progeny. This frequency is reduced for linked polymorphisms, and approaches 0% in cases of tight linkage. Additional primer sets are being developed to provide 4 - 5 markers per chromosome. For these we are using BO clones containing Tc1 elements and determining their physical map position to identify well placed polymorphisms. For those interested in trying this method we intend to make a starter kit containing each of the six primers below. reference clone in the physical map data base. We would like to thank A.Rose, M.Papp, M.Finney, D.Baillie and M.Krause for providing us with their clones containing the polymorphic sites. size of amplified fragment when primer is used with Tc1 primer 618. PCR reaction conditions: 2.5 l of worm lysis buffer containing a worm, 25 picomoles of each primer, 2.5 l 1OX standard PCR reaction buffer (100 mM Tris pH 8.3, 500 mM KCl, 15 mM MgC12, 0.01%[w/v] gelatin), 4 l dNTP mix (1.25 mM for each dNTP), 0.625 units of Taq polymerase, H20 to 25 l. 30 cycles: 94 C, 30 sec; 58 C, 60 sec; 72 C, 60 sec. Single worm PCR (from R. Barstead): Worm is picked into worm lysis buffer (50 mM KCl, 10 mM Tris pH 8.3, 2.5 mM MgC12, 0.45% NP-40, 0.45% Tween 20, 0.01% gelatin, 60 g/ml proteinase K), frozen at -70 C for 10 min, overlaid with a drop of oil, heated to 60 C 1 hr and then 95 C for 15 min.