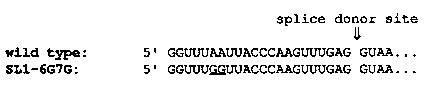Worm Breeder's Gazette 11(1): 25
These abstracts should not be cited in bibliographies. Material contained herein should be treated as personal communication and should be cited as such only with the consent of the author.
We are microinjecting altered SL1 sequences into worms in order to investigate the sequences required for splicing the C. elegans spliced leader (SL1) RNA onto the 5' end of endogenous mRNAs. The 1 kilobase pair BamHI-generated fragment encoding both the SL1 and 5S RNAs was mutagenized in vitro to produce the SL1-6G7G sequence shown [See Figure 1] A plasmid bearing the 1 kb fragment containing this altered SL1 sequence was coinjected with phage DNA containing the unc-31 gene ( generously provided by R. Hoskins) into unc-31(e928) animals. A Non- Unc line was established that segregated approximately 60% Non-Unc and 40% Unc animals per generation. Results of hybridization analysis of DNA purified from this line were consistent with the presence of lambda, plasmid and SL1-6G7G sequences. An oligonucleotide complementary to SL1-6G7G was used to probe RNA isolated from the transformed line, from Unc segregants of the transformant, and from wild-type animals. The oligonucleotide hybridized to a heterogeneous smear of RNA from the transformed line, but failed to hybridize to RNA from Unc segregants or wild type worms. This heterogeneous smear of RNA from the transformants did not hybridize to vector sequence probes or to 5S RNA probes, so it does not represent run-off transcription from the extrachromosomal DNA. These data suggest that the mutated SL1 DNA is being faithfully transcribed and spliced onto a heterogeneous collection of endogenous mRNAs. We are presently attempting to clone some of the mRNAs containing SL1-6G7G to determine if the normal SL1 specificity has been retained. We are mutagenizing SL1 further to determine the sequence and secondary structure requirements for trans-splicing in vivo.