Worm Breeder's Gazette 11(1): 21
These abstracts should not be cited in bibliographies. Material contained herein should be treated as personal communication and should be cited as such only with the consent of the author.
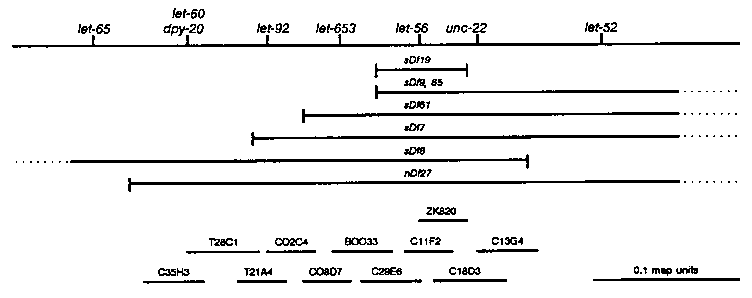
We have been studying essential genes in the unc-22 IV region of Caenorhabditis ntial genes have now been identified in this 2 map unit interval, and the minimum estimate of essential genes is 52. Lately our focus has been on the 0.2 map unit interval between dpy-20 and unc-22 (see map below). This region contains three identified essential genes with zygotic lethal mutant phenotypes. The most severe alleles of let-56, e developmental arrest in early larval stages (L1 or L2). dpy-20 (Clark and Baillie, unpublished results) and unc-22 [Moerman, Benian and Waterston, (1986) PNAS 83:2579] have both been cloned and placed on the physical map and between them lies a section of cosmid contig of about 200 kb. Coding regions on four cosmids in this interval have been mapped and characterized by identifying regions of homology with C. briggsae [Prasad and Baillie, (1989) Genomics, in press]. In order to identify the cosmids that contain essential gene coding regions, we have been injecting cosmid DNA into strains carrying a lethal mutation [let-x unc-22 nT1(IV); +/nT1(V)] and screening the F1 generation for fertile Unc-22s or Unc-22 s173) is rescued by the cosmid C11F2 and by ZK820. Stable and transient rescues were achieved, and two stable lines containing s173 and C11F2 were kept. C11F2 and ZK820 share two coding regions identified by C. briggsae homology. Two subclones containing each coding region were tried for rescue of s173 without success. Three cosmids F47H9, C29E6 and ZK822 were injected together into individuals carrying let-653(s1733) and a stable line of Unc-22 ated. Later, transient rescue using C29E6 was achieved. The stable lines are being cultured for DNA isolation so that the copy number of cosmid sequences in these lines can be examined. The map below shows the cosmids being used for transformation experiments. Note that C35H3 rescues dpy-20 but not let-60. Also note that all likely cosmids have been used in attempts to rescue let- 92(s504) and (s677), without success. This work is supported by an MDA of Canada predoctoral fellowship to DVC, and MDA of Canada and NSERC grants to DLB. [See Figure 1]