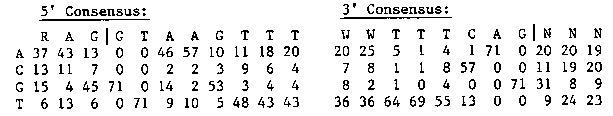Worm Breeder's Gazette 10(3): 65
These abstracts should not be cited in bibliographies. Material contained herein should be treated as personal communication and should be cited as such only with the consent of the author.
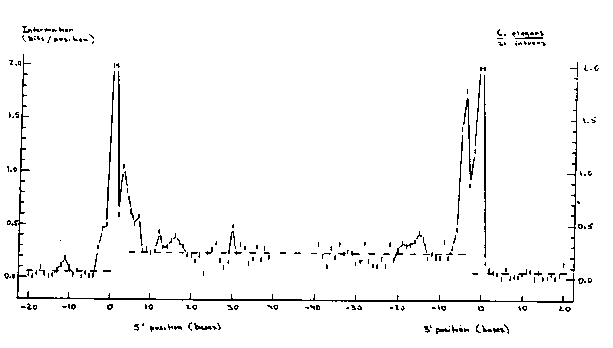
The introns of C. elegans are somewhat unusual: they are shorter than is typical for higher eukaryotes, they average roughly 74% A/T ( =W) between the splice sites, and they often lack YRAY (Y=CtT, R=A/G) lariat-formation sites (Emmons, 1988). C. elegans is, moreover, the only known higher eukaryote in which trans as well as cis splicing occurs (Krause and Hirsh, 1987; Bektesh and Hirsh, 1988; Thomas et al., 1988). These features of C. elegans introns suggest that they may encode information important for splicing at sites other than the known donor, acceptor, and lariat sites (reviewed by Sharp, 1987). A data set of 71 C. elegans introns has been collected, including 1 intron from cal-1 (Salvato et al., 1986), 4 from the hsp16 doublet ( Russnak and Candido, 1985), 8 from unc-54 (Karn et al., 1983), 4 from vit-5 (Spieth et al., 1985), 2 from vit-6 (T. Blumenthal, J. Spieth, and E. Zucker, unpublished data), 2 from col-1 and 1 from col-2 ( Kramer et al, 1982), 2 each from col-6 and col-8 and 1 each from col-7, , C. Fields, J. Kramer, B. Rosenzweig, and D. Hirsh, unpublished data), 2 each from act-1, om act-4 (M. Krause, M. Wild, and D. Hirsh, unpublished data), 7 from mec-3 (Way and Chalfie, 1988), 15 from deb-1 (R. Barstead and R. Waterston, unpublished data), 2 from dpy-13 (N. von Mende, D. Bird, P. Albert, and D. Riddle, unpublished data), and 9 from unc-22 (G. Benian, S. Nickelman, and S. Brenner, unpublished data). The donor and acceptor site consensus matrices obtained from these 71 introns are as follows: [See Figure 1] A total of 54/71, or 76% of these introns have YRAY sequences between 16 and 39 bases upstream from the conserved G of the 3' splice site; several have two or three such sequences. The information content of the 71 introns was analyzed using the method of Schneider et al. (1986); the results of this analysis are shown in Fig. 1. The two splice sites have surprisingly different structure. The 5' splice site encodes approximately 6.4 bits of information, while the 3' splice site encodes approximately 8.3 bits. The TT at -4, -5 in the 3' splice site contributes 3.2 bits, and may therefore be almost as important for identifying the splice site as the conserved AG. Satellite peaks appear on both the 5' and 3' sides of the 5' splice site. The peak at -10 corresponds to 30/71 cases of AA. The peak between 10 and 20 corresponds to T being twice as likely as A in positions 13, 18, and 19, while the peak at position 29 corresponds to a minimum (9/71) in the frequency of C/G. On the 3' side, the peak between -14 and -19 also corresponds to a minimum in the frequency of C/G. The next step in the analysis is to look for correlations between the features represented by the satellite peaks and the structures of the splice sites. Additional sequences of C. elegans introns to include in this analysis, together with the 20 bp of exonic DNA flanking each splice site, would be greatly appreciated. [See Figure 2]