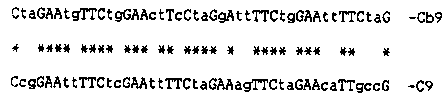Worm Breeder's Gazette 10(3): 169
These abstracts should not be cited in bibliographies. Material contained herein should be treated as personal communication and should be cited as such only with the consent of the author.
Caenorhabditis ontains a family of repetitive DNA elements, C9, that are interspersed in the genome and consist of tandemly repeated units. The tandemly repeated unit ( present in multiple copies in each element) shares the consensus (C-- GAA--TTC--G) with the 'heat-shock element' (HSE), an essential element of the heat-shock promoter in eukaryotes. The HSE is 14 base pairs long but in some heat-shock genes (2,3), two HSEs can overlap by four base pairs; this overlap is observed throughout the length of individual C9 elements. C9 members do not seem to be part of heat- shock promoters in C. elegans but a 242bp fragment of one of these elements is able to promote heat-induced transcription from a reporter gene in Saccharomyces cerevisiae (1). Seeking for clues to the events that led to the formation of the C9 family by amplification and redistribution of HSE-like sequences in the C. elegans genome, I have looked for C9-like elements in other species. Related DNA families were not detected by Southern transfer and hybridization in the genome of other eukaryotes (Saccharomyces cerevisiae, Drosophila melanogaster, Xenopus laevis) including nematodes (Dirophilaria repens, Parascaris equorum and Caenorhabditis briggsae). A feature of the C9 family is the high frequency of the restriction endonuclease XbaI recognition sites; these are found on the average every 40bp in C9 elements and extra sites can be formed every 10bp by a single base mutation without altering the HSE consensus. Genomic DNA of all the species mentioned above were digested with XbaI and end- labelled; none showed a low molecular weight, discrete DNA fraction with the sole exception of Caenorhabditis briggsae (1). I therefore performed a shot-gun experiment in which a XbaI digested, dephosphorylated plasmid DNA was mixed and ligated under appropriate conditions, with total XbaI digested genomic C. briggsae DNA. A clone with the expected features was isolated: a) it identifies, by Southern transfer, an interspersed repetitive DNA family (named Cb9) in the C. briggsae but not in the C. elegans genome; b) it contains multiple HSEs repeated in tandem with the same overlapping array present in C9. The 44 bp insert in this clone is shown below (capital letters indicate agreement with the overlapping HSE motif and asterisks indicate matches with a representative fragment from the C9 family): [See Figure 1] It is an apparent paradox that members from the two related families do not cross-hybridize, under standard experimental conditions. Probably this failure is due to the high variability of the non- conserved nucleotides in the consensus.