Worm Breeder's Gazette 10(3): 138
These abstracts should not be cited in bibliographies. Material contained herein should be treated as personal communication and should be cited as such only with the consent of the author.
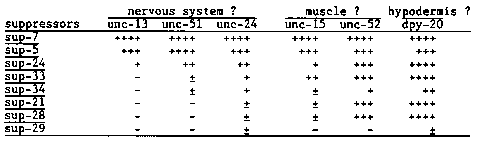
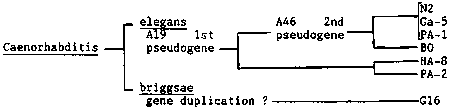
C. elegans has 12 tRNA(Trp) genes (including two probable pseudogenes), and 8 of them were converted into amber suppressors by single base changes at anticodons. We have been investigating the expression of individual members of the tRNA(Trp) gene family by cross suppression tests. Three nervous system affecting genes, unc-13(e1091, e450), unc-51(e369) and unc-24(e138), were tested. The results listed in the table indicate different suppressor activities, with the same order of suppression for the three genes. However, studies of the suppression of the two muscle-affecting genes, unc-15(e1214) ( paramyosin) and unc-52(e669) did not reveal the same order of suppressor activities found for the putative neuronal genes, suggesting tissue or developmental stage-specific expression of suppressors as discussed previously. Moreover, there is a different order of suppression between unc-15 and unc-52. This might suggest that tRNA(Trp) genes are expressed in a stage-specific manner in muscle cells, since the Unc-52 phenotype appears at a different stage than the Unc-15 phenotype. Alternatively, the unc-52 gene might be expressed in other tissue (possibly in hypodermis, because the suppression pattern of unc-52 was almost identical with that of dpy-20( e2017), which is likely to be expressed in hypodermis). [See Figure 1] We also performed a phylogenetic analysis of the tRNA(Trp) gene family in six strains of C. elegans (see figure) and a C. briggsae strain (G16). The number of the genes, RFLD (restriction fragment length difference) and two variant bases in the two genes (A19 and A46, respectively) found in N2 strains (WBG 10 #1, 54) were tested by Southern analysis using synthetic oligomers. All C. elegans strains have 12 genes, while G16 has about 13; A19 variant base was conserved in all C. elegans strains but not in G16; A46 variant base was conserved in 4 C. elegans strains but not in PA-2, HA-8 and G16; RFLDs were found in B0(2), HA-8(2), PA-2(1) and G16(no common bands with N2). The phylogeny shown in the figure was deduced. Analysis by using the tRNA(Pro) gene probe gave identical phylogeny. [See Figure 2]