Worm Breeder's Gazette 10(2): 34
These abstracts should not be cited in bibliographies. Material contained herein should be treated as personal communication and should be cited as such only with the consent of the author.
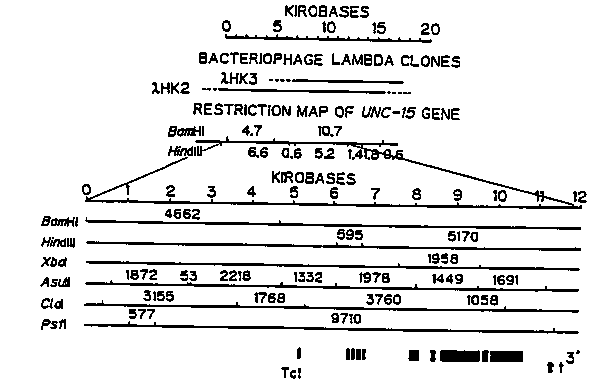
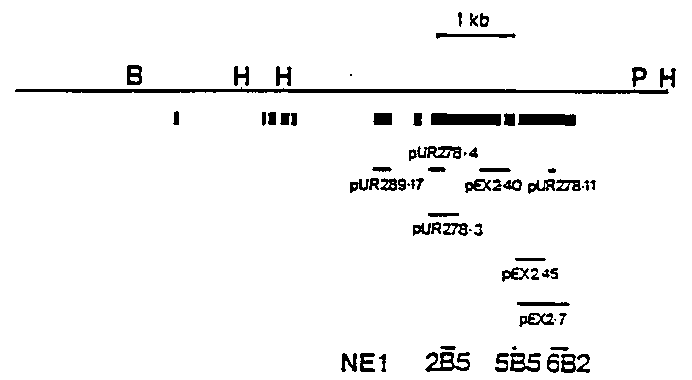
Complete genome structure of the paramyosin gene, unc15 was determined (see below). Twelve kilobases of DNA fragment from hybrid lambda HK2 were sequenced by MRC protocol of Sanger method and processed by ANALYSEQ. There were ten exons and nine introns. Consensus sequence of nine introns was GTAAGT---TTTCAG (Fig. 2). Tc1 was inserted in consensus sequence (Mori, I. et al PNAS, 85 (1988) 861-864) and was located at 196bp upstream of the initiation codon. Eight hundreds and forty amino acid residues with molecular mass of 97, 240 daltons were deduced from ten exons and the predicted amino acid sequence was completely matched to that of CNBr-fragment of the paramyosin at three different sites. The sequence was also confirmed by comparison to that of fusion protein with -galactosidase which were produced in bacterial cell and were crossreacted with anti- paramyosin serum and three monoclonal antibodies (Fig. 3). N- terminal 114 amino acid residues had 92 % homology to that of Dirofilia immitis paramyosin (Larry McReynolds, New England Biolabs). Alan Coulson sent me a letter that myo-1 contig joined to unc-15-col-7- unc-13 contig. We hope some unc-15 mutants will recover a motility with a gene therapy. [See Figures 1-3]