Worm Breeder's Gazette 10(2): 148
These abstracts should not be cited in bibliographies. Material contained herein should be treated as personal communication and should be cited as such only with the consent of the author.
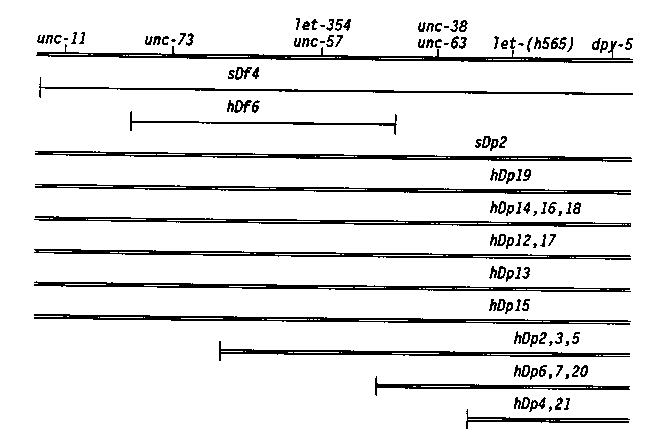
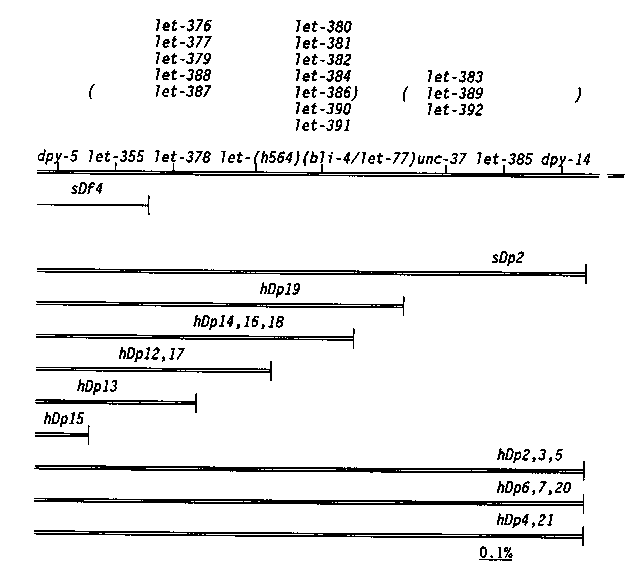
In the last newsletter we described a screen to recover derivatives of sDp2 which broke in the unc-11 Eight duplications are being used to map genes in the region. A valuable tool in this work is the translocation szT1(I;X). It has been used in the following way. unc-11 0/szT1(X) males are crossed to dpy-5 odites. All the male progeny are unc-11 dpy-5 of the of the I;X pseudolinkage in szT1 strains. When these males are crossed to unc-11 unc-11 -11-) strains, all the wild-type cross progeny are unc-11 unc-x egregants from these wild types are observed to determine if the hDpz covers the unc-x mutation. In this manner, the duplication breakpoints shown below were determined. [See Figure 1] In a similar screen, we have isolated derivatives of sDp2 which break between dpy-5 and dpy-14. Phenotypically wild type dpy-5 dpy-5 sDp2 worms were treated with 1500 or 3000 R and rare Dpy worms were recovered. From 7000 F1s screened, 10 exceptional Dpys were recovered. One isolate was a dominant Dpy and another a recessive Dpy mutation. The remaining eight were duplications which cover dpy-5 but not dpy-14.These duplications are being used to map essential genes. All the essential genes rescued by the duplication sDp2, reported in Howell et al. ( Genet. Res. 49:207) as mapping to the right of dpy-5 have been mapped using the duplication hDp19. Four of these 17 essential loci were outside (to the right of) the hDp19 breakpoint. This data places the breakpoint of hDp19 12 map units to the right of dpy-5 (three- quarters of the distance between dpy-5 and dpy-14). One of the duplications, hDp15, does not cover let-355, which maps in sDf4, but previously was inseparable from dpy-5. This result places the sDf4 breakpoint to the right of dpy-5 (this was not previously known because sDf4 was induced on a dpy-5 chromosome). [See Figure 2] The 16 duplications and two deficiencies (sDf4; potential to divide the 4 map unit interval between unc-11 regions. These and additional duplications which are being generated will be used to assist in the identification and mapping of essential genes in this region.