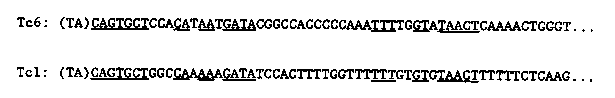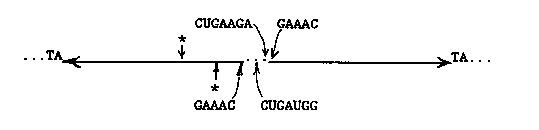Worm Breeder's Gazette 10(2): 145
These abstracts should not be cited in bibliographies. Material contained herein should be treated as personal communication and should be cited as such only with the consent of the author.
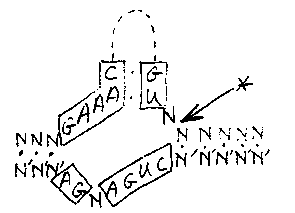
We are continuing to study a family of palindromic sequences initially identified as the cause of a 1.6 kb Bergerac (filled site) /Bristol (empty site) DNA polymorphism. The foldback DNA sequence inserted in Bergerac at the polymorphic site we now call Tc6. We are currently cloning several new Tc6-homologous restriction fragments that are present in genomic Southern hybridizations of DNA from mutator strains TR679 and RW7406. We wish to determine whether these are caused by recent Tc6 transposition events. Tc6 does not cross- hybridize with Tc1, Tc2, Tc3, Tc4, or Tc5. We have previously reported that the 7 nucleotides at the ends of Tc6 are identical to the terminal nucleotides of Tc1, and that Tc6 elements (three out of three examined), like Tc1 elements, are inserted at TA dinucleotides. We now point out additional similarities between the two transposons. As shown in Figure 1, two additional runs of sequence similarity are found, identically positioned with respect to the ends of the two transposons. Computer searches uncover no additional significant similarities. [See Figure 1] Because the palindromic structure of Tc6 is similar to the 'rod' forms of the RNA genomes of plant-pathogenic viroids and virusoids, and because short runs of sequence similarity to these agents were also present in Tc6, we carried out a careful comparison of Tc6 and viroid and virusoid sequences. We found in both strands of Tc6 sequences that could form a structure known as a 'hammerhead' (Figure 2), which can be formed from the sequences of all known viroids and virusoids (Forster and Symons, Cell 49, 211-220, 1987; Uhlenbeck, Nature 328, 596-600, 1987), as well as from the self-cleaving transcripts of a newt satellite (Epstein and Gall, Cell 48, 535-543, 1987). All these RNA's undergo self-catalyzed cleavage of the phosphodiester backbone within the 'hammerhead' at the site indicated by the arrow in Figure 2. The Tc6 hammerheads contain nearly all of the absolutely conserved nucleotides of the consensus structure (boxed in Figure 2). The conserved CUGANGA sequences are within the 75 nucleotide non-repeated central region, the conserved GAAAC sequences are just inside the palindromic arms, and the cleavage sites predicted by the hammerhead structures are in one of the arms (figure 3). [See Figure 2] We therefore examined the properties of transcripts of Tc6 synthesized in vitro. We find that transcripts of both strands cleave in the transcription buffer with nearly 100% efficiency. The sites of cleavage, determined from the mobilities of the products on gels, are consistent with the predicted cleavage sites in the hypothetical hammerhead structures. The fact that both strands cleave within the palindromic arms, but at different sites, is consistent with a self catalyzed cleavage directed by the secondary structure of each RNA molecule. [See Figure 3] In the case of viroids and virusoids, RNA self-cleavage is thought to play a role in generating monomer genomes from concatemeric, rolling circle replication intermediates. Significantly, at one genomic site in C. elegans we find a tandem array of possibly five 1. 6 kb Tc6 elements, reminiscent of such concatemeric intermediates. Taken altogether, these data suggest that Tc6 replicates in the form of an RNA intermediate. Transposition could result from reverse transcription of circular monomers or multimers produced by self- cleavage and ligation, followed by insertion of DNA copies into the genome by a transposase used in common with Tc1. It is interesting to note that Uhlenbeck (op cit) has shown that an oligonucleotide of the sequence shown for the bottom strand in Figure 2 will catalytically cleave in trans an oligonucleotide with the sequence of the upper strand. The lower oligonucleotide is therefore a sequence-specific ribonuclease, with its specificity determined by the sequences indicated by N'. This suggests the possibility of synthesizing a ribonucleolytic RNA with specificity for a sequence in the mRNA of a gene whose function one wishes to eliminate, and expressing this RNA in vivo from an inducible promoter.