Worm Breeder's Gazette 10(2): 140
These abstracts should not be cited in bibliographies. Material contained herein should be treated as personal communication and should be cited as such only with the consent of the author.
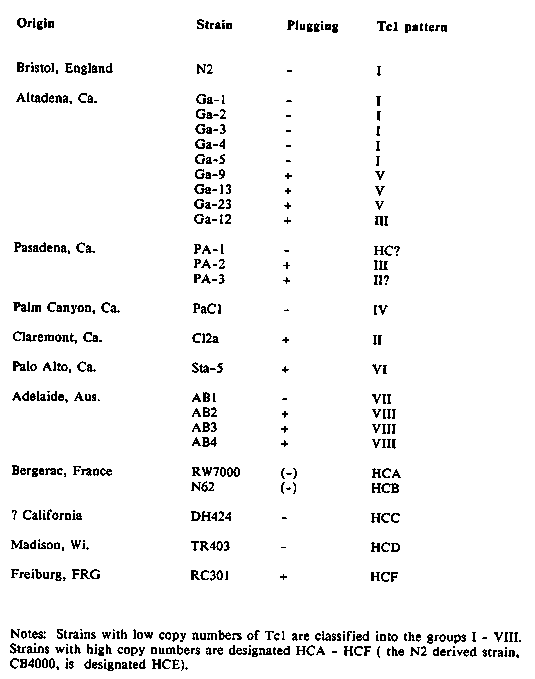
Many natural isolates of C. elegans differ from the standard laboratory strain, N2, in copulatory plug formation: the males from these strains deposit a gelatinous blob over the vulva of a mated hermaphrodite, but N2 males do not. Previously (Hodgkin, Doniach & Kenyon, WBG 8-3:36) we showed that the plugging trait, as discovered in the wild isolate Sta-5 (formerly named Cal-5) behaved as a Mendelian dominant, mapping to the locus plg-1 on LGIII. We suggested that N2 had lost the plugging trait at some point during its scientific career. A survey of natural isolates indicates that this is not the case; instead, it appears that many natural races are nonpluggers. Male stocks were established from the 24 strains listed in the table, and males were tested for plug formation. All non- plugging strains were crossed with N2, and F1 heterozygous males tested: all were negative. Thus, if non-plugging represents loss-of- function, then all these strains carry recessive mutations of the same gene, plg-1. Also, the plugging trait in two of the positive strains, AB3 and RC301, was examined genetically, by crossing with N2-derived mapping strains. In both cases the trait behaved as a dominant mapping to approximately the same location as plg-1.In almost all of these races (or at least the stocks of them presently held in Cambridge) males are present at less than 1% of the individuals in a growing population, but these males are always potent and male stocks are easily established. The only exceptions are the two Bergerac strains, RW7000 and N62, which are well known to produce males that mate very poorly. It is not obvious what advantage or disadvantage is associated with plugging. The trait is stable in culture: both plugging and non- plugging strains maintain their character over hundreds of generations. DNA from these strains has been examined using transposon probes. As previously reported by others (Katzenberg & Emmons, WBG 7-2:34; Riddle et al., WBG 8-2:52), the pattern of Tc1-hybridizing bands in a restriction digest can be used to classify the different races. A Tc3 probe can also be used, and gives similar results, where tested. The data confirm (with one anomaly) and extend the classification of Katzenberg and Emmons, which is therefore used in the table. The anomaly is our strain of 'PA1', which may have mutated into a high- copy strain, but this is a preliminary result. Note that for strains from the same locale, the plugging status and the racial classification are concordant. Use of other probes, or different digests, might however subdivide some of the races. As other workers have noticed (e.g. Cassada, WBG 9-3:29), many of the different races exhibit slight behavioral and/or morphological differences from N2. It is likely that they could be used as good sources of natural variability for experiments in population genetics, evolution, and so on. Also, in view of the general intention to learn as much as possible about the biology of C. elegans, it remains desirable to obtain further natural isolates of this species, especially from exotic parts of the world. I am grateful to Phil Anderson, Randy Cassada, Tabby Doniach, Ed Hedgecock, David Hirsh, Carl Johnson, Don Riddle and Dick Russell for collecting and providing the strains examined. [See Figure 1]