Worm Breeder's Gazette 10(2): 120
These abstracts should not be cited in bibliographies. Material contained herein should be treated as personal communication and should be cited as such only with the consent of the author.
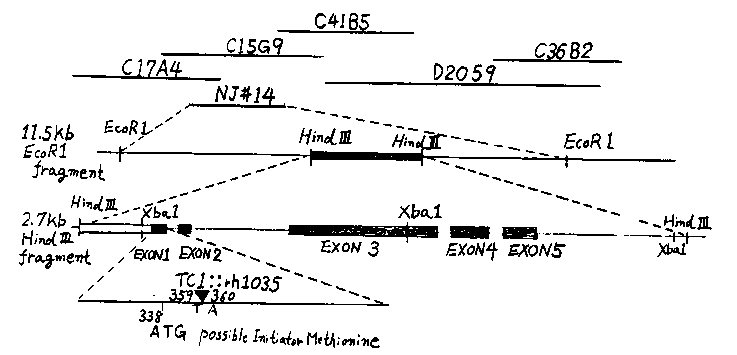
In the previous issue (WBG 10(1) p.35), we reported cloning a small DNA fragment containing the Tc1 insertion which caused the unc-6 ( rh1035) mutation. Since then, we have isolated an 11.5 kb EcoRI fragment including this region from N2 DNA (NJ#14, EMBL4 vector) Alan Coulson has kindly positioned this fragment on the MRC genome map. We have now sequenced both the original 1.25 kb XbaI fragment of unc-6( rh1035) DNA (plasmid NJ#6) and a 2.7 kb HindIII fragment of N2 DNA subcloned from phage NJ#14. The 2.7 kb HindIII fragment has five exons which encode a hypothetical protein fragment of 360 residues. This fragment is not closely related to any protein in the current PIR database The last exon in the 2.7 kb HindIII fragment is followed by an intron which continues beyond the fragment. The first exon is preceded by a non-coding region which may be part of an intron or the unc-6 promoter region. In fact, a possible 21 residue signal sequence with an initiator methionine occurs in the first exon shortly upstream of the Tc1::rh1035 insertion site. TATA and CCAAT motifs occur about 90 and 120 bp upstream of the proposed initiator codon. The decanucleotide sequence TTTGCCTCTC is tandemly repeated about 200 bp upstream of this codon. [See Figure 1]