Worm Breeder's Gazette 10(2): 111
These abstracts should not be cited in bibliographies. Material contained herein should be treated as personal communication and should be cited as such only with the consent of the author.
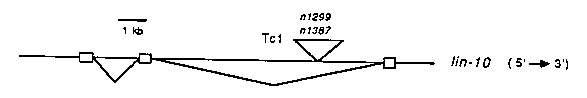
Each of the six Pn.p cells in the vulval equivalence group (P3.p to P8.p) has the same developmental potential. These cells can undergo any one of three cell lineages depending primarily on the proximity of a regulatory cell, the anchor cell. When the anchor cell is near, Pn. p cells undergo vulval cell lineages (termed the 1 and 2 cell lineages), and when the anchor cell is distant, they undergo a non- vulval hypodermal cell lineage (termed the 3 cell lineage). lin-10 is involved in specifying Pn.p cell lineages; mutations in this gene cause all Pn.p cells to express the 3 cell lineage, resulting in a Vulvaless (Vul) phenotype. lin-10 probably acts in Pn.p cells and thus may be involved in signal reception, signal transduction or in the control of gene expression. Mutations in most genes involved in Pn.p cell determination have pleiotropic effects, and in at least some cases, this is because these genes control other cell lineages. To see if lin-10 is also involved in specifying the fates of non-vulval cells, we wanted to determine its null phenotype. Our results suggest that lin-10 function is required only in the Pn.p cells, since a complete lack of lin-10 gene activity results in a Vul phenotype. First, ten lin-10 alleles have been isolated by screening for Vul mutants to date. Thus, Vul mutations in lin-10 arise about as frequently as mutations in any gene involved in Pn.p cell determination and are isolated more frequently than alleles of genes such as lin-3 and let-23, which have a lethal null phenotype. Second, the Vul phenotype is not enhanced when lin-10( e1439) is placed in trans to a deficiency. Third, we isolated eight lin-10 alleles by screening for mutations that failed to complement lin-10(e1439). One allele, n1638, appears to be amber-suppressible. All of these alleles result in a Vul phenotype when homozygous, with no obvious pleiotropic effects. Thus, lin-10 appears to control specifically the fates of only three cells: P5.p, P6.p and P7.p. Using the approach of transposon tagging, we have cloned a gene that is probably lin-10 (see figure below and CSH abstracts, 1987). In two mut-2-induced a Tc1 element has inserted into a 10 kb intron of a candidate lin-10 transcription unit. This gene produces a 1.6 kb mRNA, and its expression is reduced both in the transposon insertion alleles and in an EMS-induced lin-10 point mutant, n299. These results suggest that this transcription unit corresponds to the lin-10 gene. Based upon preliminary microinjection experiments, a cosmid containing this gene appears able to rescue the Vul phenotype of a lin-10 strain. [See Figure 1] We have isolated three full-length lin-10 cDNA clones. Sequence analysis of these and other cDNA clones shows that this gene encodes a protein (407 amino acids) that is not homologous to sequences in any of the protein databases (NBRF/PIR Protein Database including the prerelease sequences and the Pasteur Institute Protein Database) nor is it striking in its primary structure. To determine when and where lin-10 is expressed, we have isolated anti-lin-10 antisera. When these antisera are used to probe Western blots, one major band (45 kd) is observed in samples derived from N2, and this band is reduced in intensity in samples derived from either of the transposon-insertion alleles or from a point mutant, n299. This result suggests that these antisera are specific for lin-10. The anti-lin-10 antisera were then used to stain worms in situ. Embryos stain the strongest, from the single cell stage to just before hatching. This staining is reduced in a strain carrying lin-10(n1299), a transposon insertion allele. The embryonic expression of this gene is considerably earlier than the time of vulva formation, which is in the third larval stage. One possibility is that the low level of expression during the third larval stage is sufficient for vulval determination. Another possibility is that lin-10 acts early in development, endowing cells with the potential to undergo vulval induction at some later time. These possibilities might be distinguished by determining the temperature-sensitive period of lin- 10(n1509ts). The anti-lin-10 antisera detect an antigen localized to the cytoplasm, suggesting that lin-10 does not bind DNA nor does it function as an extracellular receptor. Staining with anti-lin-10 antisera is observed in most or all cells in the embryo. There are a variety of genes that, like lin-10, display a specific mutant phenotype but are expressed more generally ( e.g. the human retinoblastoma oncogene). Such a pattern of expression can be rationalized in two general ways. First, at the cellular level, lin-10 is similar to genes with wild-type null phenotypes (e.g. unc-93 and act-3), since in cells in which lin-10 is expressed the loss of lin-10 activity has no effect. Perhaps in these cells, but not in the Pn.p cells, there is an activity that is redundant with lin-10. Second, although lin-10 is expressed in many embryonic cells, it might function only in Pn.p cells. For example, lin-10 might be regulated at the posttranslational level so that active gene product appears in only P5.p, P6.p and P7.p. Alternatively, lin-10 may play a role in signal transduction and function only in those cells that actually receive a signal. Finally, since Pn.p cell determination is controlled by a genetic pathway containing at least seventeen genes, the expression of only some of the genes in this pathway may be specific to Pn.p cells. As a result, lin-10 may have functional consequences only in Pn.p cells because only in these cells are all of the genes of the lin-10 pathway expressed.