Worm Breeder's Gazette 10(2): 110
These abstracts should not be cited in bibliographies. Material contained herein should be treated as personal communication and should be cited as such only with the consent of the author.
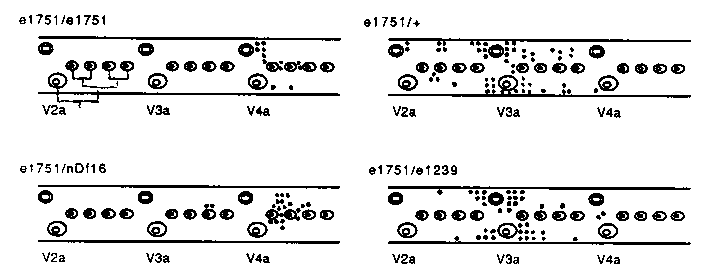
QL and QR are generated through identical patterns of division from ABpl and ABpr, and they divide identically, to produce three neurons and two programmed cell deaths. Yet QL descendants stay near V5.a or migrate back into the tail, while QR and its descendants migrate anteriorly past the gonad into the vicinity of V2.a or the head. Five genes are known to influence the direction of migration of these neuroblasts. Recessive mutations in the genes mab-5, 3), and hch-1 cause both QL and QR to migrate anteriorly, while a semidominant mutation in the gene lin-21 prevents QR and its descendants from migrating past the gonad. There are three reasons to suspect that lin-21(e1751) might be a gain of function allele of mab-5. First, e1751 has the opposite effect on Q migration. Second, mab-5 and lin-21 have not been separated in mapping experiments. 16 recombination events between dpy- 17(e164) and unc-32(e189) failed to separate mab-5(e1239) and lin-21( e1751) (E. Hedgecock, personal communication). Third, a restriction fragment length polymorphism associated with e1751 has been seen on two Southern blots probed with putative mab-5 DNA sequences. (R. Hoskins, MRC pers. comm., repeated by M. Costa, UCSF) If mab-5(e1239) is a null allele of lin-21, then it should behave like a deficiency when in trans to e1751. Otherwise, since known mab- 5 alleles are recessive, e1751/e1239 should resemble e1751/+. Here are the positions of Q descendents in L2 heterozygotes. There were regularly two Q descendants in the body region illustrated: [See Figure 1] First, since e1751/nDf16= e1751/e1751, e1751/+, e1751 is likely to be a neomorphic mutation whose effects on Q migration can be alleviated by wild type gene activity. Second, the phenotype of e1751/e1239 resembles that of e1751/+, while the phenotype of e1751/nDf16 is quite different, and resembles the e1751/e1751 phenotype. This suggests, as argued above, that e1751 is not an allele of mab-5. However, it is always possible that e1239, while null for the mab-5 wild-type activities, retains activity in suppressing e1751. Examination of more mab-5 alleles over e1751 will help rule out this possibility.