Worm Breeder's Gazette 10(1): 96
These abstracts should not be cited in bibliographies. Material contained herein should be treated as personal communication and should be cited as such only with the consent of the author.
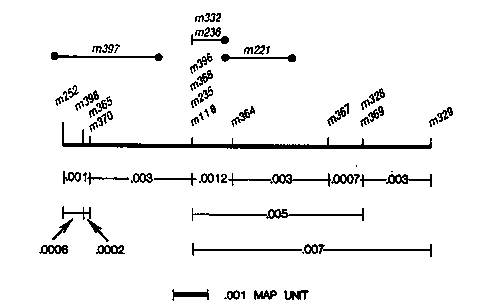
A fine-structure genetic map has been constructed for ama-1, an essential gene encoding the amanitin-binding subunit of RNA polymerase II. Sixteen EMS-induced recessive lethal mutations have been positioned in the gene by determining their intragenic recombination distances from m118, a mutation that confers dominant resistance to alpha-amanitin. The 16 mutants, all isolated in the ama-1(m118) background, include 13 that are early larval lethals, and three that are mid-larval lethals at 25 C. Six of the mutants exhibit temperature-dependence in the severity of their phenotype. Intragenic recombination between the lethal site and the parental resistance mutation was detected by means of resistance to amanitin. Recombinants were detected at frequencies as low as 2x10+E-6. The segregation of closely linked flanking markers, unc17 and unc-5, revealed whether the mutation was to the left or the right of m118. By adding the distances between the extreme left and right mutations, the ama-1 gene is estimated to be 0.011 map units long, with m118 positioned 0.004 map units from the left-most mutation. To order the lethal mutations with respect to each other, viable heteroallelic strains were constructed using the free duplication, mDp1[unc17(e113) dpy-13(+)ama-1(f)(+). The heteroallelic strains were sensitive to amanitin, and recombination events between the lethal mutations were specifically selected by means of the dominant amanitin resistance encoded on the recombinant chromosome. The segregation of outside markers revealed the order of the lethal mutations. The ama-1 fine- structure map is linear, and additive, based on data obtained from mapping lethal alleles relative to m118, and on data obtained from mapping lethal alleles relative to each other (Figure 1). The distribution of mutations within the gene is non-random. The early larval lethal mutations map throughout the gene. Arrested development and death in the L1 stage is the ama-1 null phenotype. Evidence for this comes from the mDf10 deficiency, which conveys an L1- lethal phenotype in deficiency homozygotes (Rogalski and Riddle, Genetics, in press). Five mutations are clustered very close to the resistance mutation, m118, which maps approximately 1/3 of the way through the gene from the dpy-13 (left) end. Molecular analysis has shown that the dpy-13 end is the 5' end of the gene (D. Bird and D. Riddle, unpublished). Six lethal mutations are positioned to the left of m118, and six are to the right of the m118 cluster. Hence, the density of mutations is twice as great near the left end of the gene ( encoding the N-terminus of the protein) than it is on the right side. The amanitin-binding site is highly conserved among divergent species, and this region might be essential to the structural integrity of the large subunit. Attempts to purify RNA polymerase II from strains heterozygous for the sterile mutations m236 and m235 revealed amanitin-resistant species that were degraded rapidly during the first steps of purification (M. Golomb, personal communication). It is interesting to note that four of the five sterile mutations ( m235, m368, m369, m236, and m396) map very close to m118, also suggesting that this region of the large subunit is important in enzyme stability. These mutants all exhibit temperature-sensitive phenotypes (larval lethal at 25 C). [See Figure 1]