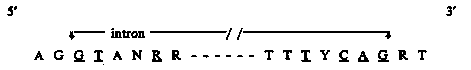Worm Breeder's Gazette 10(1): 55
These abstracts should not be cited in bibliographies. Material contained herein should be treated as personal communication and should be cited as such only with the consent of the author.
We have sequenced approximately 8.7 kb of genomic DNA, implicated by transcript mapping experiments as spanning ama-1. Removal of sequences resembling introns leaves a sequence which translates to a continuous open reading frame of 1795 amino acids. Alignment of this deduced sequence with that encoding the large subunit of yeast RNA polymerase II (RP02 1 ) indicates that ama-1. encodes the analogous protein in C. elegans, and that our sequence begins approximately at the codon for amino acid 20. Currently, we are cloning adjacent fragments so as to complete the ama-1 sequence. We have, however, made some preliminary observations. The ama-1 gene is broken into 11 exons, ranging in size from 109 bp to 1589 bp. The introns generally are large by worm standards. Most are around 300 bp; the largest is 462 bp and the smallest is 49 bp. The splice junctions have the consensus: (with underlined bases conserved). [See Figure 1] The length of the 3' untranslated region may be extremely short. The only appropriate AAUAAA motif is immediately adjacent to the stop codon, with a TGTGT box 34 bp further 3'. We plan to confirm the location of the 3' terminus experimentally. The homology of the deduced amino acid sequence of ama-1 with that of yeast RP021 extends along the whole length of both peptides. The identity (Wilbur and Lipman algorithm) is approximately 50% overall, but includes several regions of more than 30 amino acids common to both polypeptides. The carboxy terminus of ama-1 is composed of a 'tail' of 31 tandem repeats of a heptapeptide with consensus: tyr, ser, pro, thr, ser, pro, ser. A similar tail is present in the yeast (26 repeats) and mouse ( 52 repeats) polypeptides. In addition to sequencing the wild-type gene, we have sequenced a portion of m118, an EMS-induced alpha-amanitin resistance allele of ama-1. We have detected a single GC->AT change, resulting in a cys -> tyr substitution. When the amino acid sequences are aligned to maximize homology, this change falls at a position only 14 bp from the site of the base which is altered in a mouse X-amanitin resistant mutant cell line (Bartolomei and Corden, Mol. Cell. Biol. 7: 586- 594, 1987). This position is consistent with the genetic fine- structure map position of the m118 mutation. We are planning a functional test to confirm that this sequence change indeed conveys resistance to alpha-amanitin.