Worm Breeder's Gazette 10(1): 54
These abstracts should not be cited in bibliographies. Material contained herein should be treated as personal communication and should be cited as such only with the consent of the author.
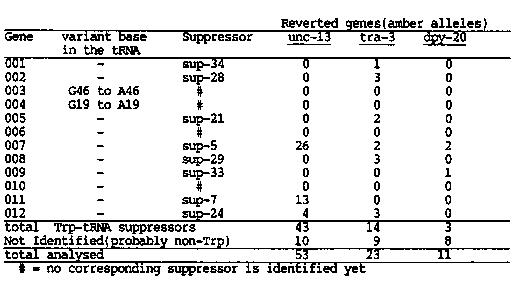
C. elegans has 12 tRNA(Trp)(UAG) genes. We have cloned and sequenced all of them. Preliminary sequencing data indicate 10 out of 12 members encode identical tRNAs while 2 of them carry single base changes at highly conserved positions; G19 to A19 and G46 to A46, respectively. Flanking sequences show only limited homology. We have made synthetic oligomers complementary to the anticodon region of either wild type tRNA(Trp)(UAG)or amber suppressor tRNA(Trp)(UAG). Each oligomer (19-mer) was [32P]-labeled and used as a probe for genomic Southern blot to look for the one base change of the tRNA(Trp) genes. Only the corresponding gene lights up under our hybridization conditions. From the sequencing results, this should reveal any change in 12 tRNA(Trp) genes at the anticodon region and would overcome the shortcoming of the previous XbaI Southern analysis where not all the tRNA(Trp) genes are resolved. We analyzed the previously unidentified class of unc-13 and tra-3 revertants, and 14 more revertants of tra-3 and 11 dpy-20 revertants. The unc-13 revertants contained no new tRNA(Trp) suppressors. But two of the tra-3 suppressors, sup-21(e1957) and sup-21(e2064), turned out to be tRNA( Trp)(UAG) suppressors of the same gene which escaped detection in the XbaI Southern analysis. And one of the new tra-3 suppressors was a new gene of tRNA(Trp)(UAG) 34(e2227). Among 11 dpy-20 revertants, two were due to sup-5 and one suppressor was identified as a tRNA(Trp) amber suppressor another new gene, sup-33(st389). 8 out of 12 members of tRNA(Trp) genes have been converted into amber suppressors so far. Two of the remaining might be pseudogenes and two are left. We are testing activities of these new suppressors. Our results to date are summarized below. [See Figure 1]