Worm Breeder's Gazette 10(1): 29
These abstracts should not be cited in bibliographies. Material contained herein should be treated as personal communication and should be cited as such only with the consent of the author.
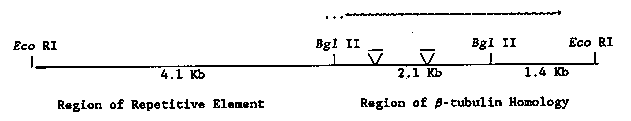
Benomyl and other benzimidazole carbamates interfere with the polymerization of microtubules. In the presence of benomyl, wild-type C. elegans grows slowly, is severely uncoordinated, and has fewer neuronal processes in the ventral cord. Mutations conferring resistance to this drug map to one gene, ben-1, on chromosome III. Virtually all of the EMS-induced and the TR679-derived alleles result in a temperature-dependent dominant drug resistance. ben-1 animals have no visible phenotype when grown in the absence of benomyl. Since benomyl resistance is conferred by mutations in -tubulin genes in Aspergillus, Saccharomyces, and Physarum, we have performed low stringency hybridizations of -tubulin sequences (provided by Linda Gremke and Joe Culotti) to DNA isolated from ben-1 mutants. DNA from two of the TR679 derived strains contains an insertion of approximately 2.1 kb in one of the -tubulin homologous restriction fragments. We have cloned a 7.6 EcoRI fragment from wild-type C. elegans that corresponds to this -tubulin. The -tubulin homology maps to the right end of the fragment, as depicted below. The left end of this fragment contains a repetitive DNA element (a non-Tc1 homologous sequence present in approximately 30 copies in C. elegans). The two transposon insertions are localized to different sites in the 2.1 kb BglII fragment. Although these are clearly not Tc1 insertions, we have not yet identified which, if any, of the characterized transposons are associated with the ben-1 mutations. Preliminary transcription analysis suggests that the ben-1 mRNA is abundant in wild-type animals. (Consistent with this observation, we have been successful in isolating a highly homologous clone from Julie Ahringer's cDNA library.) The transcript is approximately 3 kb and the direction of transcription is from left to right as shown. Molecular analysis has provided some insights into the mechanism of benomyl resistance. One of our ben-1 alleles (derived from TR679) is actually a deletion of the 2.1 and 1.4 kb restriction fragments. Since this deletion allele is dominant, it appears that a critical level of ben-1 gene product is required for sensitivity to benomyl. Genetic and molecular experiments directed towards confirming this hypothesis are underway.