Worm Breeder's Gazette 10(1): 18
These abstracts should not be cited in bibliographies. Material contained herein should be treated as personal communication and should be cited as such only with the consent of the author.
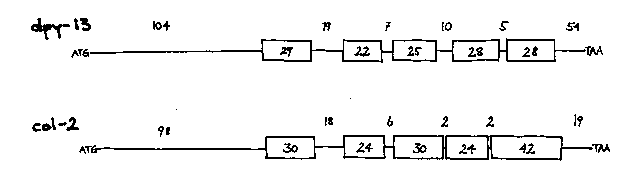
The dpy-13 gene is located 0.05 map units to the left of ama-1 IV, a gene that has been cloned and sequenced (see Bird and Riddle, this issue). As we reported at the C. elegans Meeting last May, we isolated three spontaneous dpy-13 mutants in Tc1 'high-hopper' strains (given to us by Moerman and Waterston) then used cosmid sequences ( provided by Sulston and Coulson) to identify a small region 10 kb from ama-1 that carried a 1.6 kb Tc1 insert in each of the three mutants, but lacked the insert in two revertants tested. Hybridization of the dpy-13 clone, a 1.25 kb HindIII/EcoRI fragment, to the genomic DNA from the EMS-induced dpy-13 mutants, e184 and e458, revealed deletions of about 30 bp and 250 bp, respectively. The left endpoint of a deficiency, mDf4, that included ama-1 was known from genetic analysis to be nearby, and this was located 8.2 kb to the left of dpy-13.On a northern blot with wild-type RNA, the dpy-13 clone detected a transcript at 1.1 kb. An additional band of 1.25 kb was detected in RNA from one of the Tc1 insertion mutants. The 1.25 kb species may represent missplicing resulting from the Tc1 insert. The normal size 1.1 kb transcript also found in the dpy-13::Tc1 could represent alternative splicing, or it might be the transcript of a related gene. Even at high stringency, the 1.25 kb probe hybridizes weakly to more than a dozen fragments in digests of genomic DNA. This homology extends throughout the fragment. The Tc1 insertion sites were found to be clustered near the HindIII site, and DNA sequencing revealed that this is the 5' end of the gene. Nearly all of the gene has now been sequenced. Including three short (~50 bp) introns, the distance from the initiator methionine codon to the stop codon is 1.15 kb. Transcription is from left to right on the genetic map. The DNA sequence reveals an even closer relationship to collagen genes than we reported at the C. elegans Meeting. There is a general similarity in the deduced amino acid sequence (Figure 1), especially to col-2 (Kramer et al., 1982, Cell 30: 599-606). Both genes encode five short helical regions with glycine as every third amino acid. When hybridized to genomic DNA at moderate stringency, both dpy-13 and col-2 detect the same bands, which presumably represent a subset of collagen genes in C. elegans. The region around ama-1 appears to be rich in collagen genes; ama-1 is flanked by two such genes that have not been identified genetically. The dpy-13 gene, however, seems to have a unique function, since recessive mutations affecting body length are detectable in this gene.